High-bandwidth protein analysis using solid-state nanopores
- PMID: 24507610
- PMCID: PMC3944622
- DOI: 10.1016/j.bpj.2013.12.025
High-bandwidth protein analysis using solid-state nanopores
Abstract
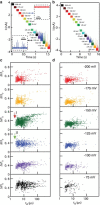
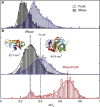
High-bandwidth measurements of the ion current through hafnium oxide and silicon nitride nanopores allow the analysis of sub-30 kD protein molecules with unprecedented time resolution and detection efficiency. Measured capture rates suggest that at moderate transmembrane bias values, a substantial fraction of protein translocation events are detected. Our dwell-time resolution of 2.5 μs enables translocation time distributions to be fit to a first-passage time distribution derived from a 1D diffusion-drift model. The fits yield drift velocities that scale linearly with voltage, consistent with an electrophoretic process. Further, protein diffusion constants (D) are lower than the bulk diffusion constants (D0) by a factor of ~50, and are voltage-independent in the regime tested. We reason that deviations of D from D0 are a result of confinement-driven pore/protein interactions, previously observed in porous systems. A straightforward Kramers model for this inhibited diffusion points to 9- to 12-kJ/mol interactions of the proteins with the nanopore. Reduction of μ and D are found to be material-dependent. Comparison of current-blockage levels of each protein yields volumetric information for the two proteins that is in good agreement with dynamic light scattering measurements. Finally, detection of a protein-protein complex is achieved.
Copyright © 2014 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Figures




References
-
- Movileanu L. Single-molecule detection of proteins using nanopores. In: Barth F.G., Humphrey J.A.C., Srinivasan M.V., editors. Frontiers in Sensing: From Biology to Engineering. Springer-Verlag; Vienna, Austria: 2012. pp. 363–381.
-
- Gu L.Q., Braha O., Bayley H. Stochastic sensing of organic analytes by a pore-forming protein containing a molecular adapter. Nature. 1999;398:686–690. - PubMed
-
- Xie H.Z., Braha O., Bayley H. Single-molecule observation of the catalytic subunit of cAMP-dependent protein kinase binding to an inhibitor peptide. Chem. Biol. 2005;12:109–120. - PubMed
-
- Siwy Z., Trofin L., Martin C.R. Protein biosensors based on biofunctionalized conical gold nanotubes. J. Am. Chem. Soc. 2005;127:5000–5001. - PubMed
-
- Sexton L.T., Horne L.P., Martin C.R. Resistive-pulse studies of proteins and protein/antibody complexes using a conical nanotube sensor. J. Am. Chem. Soc. 2007;129:13144–13152. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

