Autophagy-dependent production of secreted factors facilitates oncogenic RAS-driven invasion
- PMID: 24513958
- PMCID: PMC3980002
- DOI: 10.1158/2159-8290.CD-13-0841
Autophagy-dependent production of secreted factors facilitates oncogenic RAS-driven invasion
Abstract
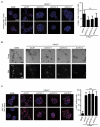
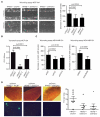
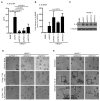
The tumor-promoting functions of autophagy are primarily attributed to its ability to promote cancer cell survival. However, emerging evidence suggests that autophagy plays other roles during tumorigenesis. Here, we uncover that autophagy promotes oncogenic RAS-driven invasion. In epithelial cells transformed with oncogenic RAS, depletion of autophagy-related genes suppresses invasion in three-dimensional culture, decreases cell motility, and reduces pulmonary metastases in vivo. Treatment with conditioned media from autophagy-competent cells rescues the invasive capacity of autophagy-deficient cells, indicating that these cells fail to secrete factors required for RAS-driven invasion. Reduced autophagy diminishes the secretion of the promigratory cytokine interleukin-6 (IL-6), which is necessary to restore invasion of autophagy-deficient cells. Moreover, autophagy-deficient cells exhibit reduced levels of matrix metalloproteinase 2 and WNT5A. These results support a previously unrecognized function for autophagy in promoting cancer cell invasion via the coordinate production of multiple secreted factors.
Significance: Our results delineate a previously unrecognized function for autophagy in facilitating oncogenic RAS-driven invasion. We demonstrate that an intact autophagy pathway is required for the elaboration of multiple secreted factors favoring invasion, including IL-6.
Figures




References
-
- Schubbert S, Shannon K, Bollag G. Hyperactive Ras in developmental disorders and cancer. Nat Rev Cancer. 2007;7:295–308. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

