Identification of novel, highly expressed retroviral microRNAs in cells infected by bovine foamy virus
- PMID: 24522910
- PMCID: PMC3993790
- DOI: 10.1128/JVI.03587-13
Identification of novel, highly expressed retroviral microRNAs in cells infected by bovine foamy virus
Abstract
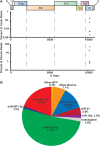
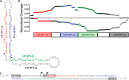
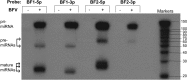
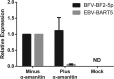
While numerous viral microRNAs (miRNAs) expressed by DNA viruses, especially herpesvirus family members, have been reported, there have been very few reports of miRNAs derived from RNA viruses. Here we describe three miRNAs expressed by bovine foamy virus (BFV), a member of the spumavirus subfamily of retroviruses, in both BFV-infected cultured cells and BFV-infected cattle. All three viral miRNAs are initially expressed in the form of an ∼ 122-nucleotide (nt) pri-miRNA, encoded within the BFV long terminal repeat U3 region, that is subsequently cleaved to generate two pre-miRNAs that are then processed to yield three distinct, biologically active miRNAs. The BFV pri-miRNA is transcribed by RNA polymerase III, and the three resultant mature miRNAs were found to contribute a remarkable ∼ 70% of all miRNAs expressed in BFV-infected cells. These data document the second example of a retrovirus that is able to express viral miRNAs by using embedded proviral RNA polymerase III promoters.
Importance: Foamy viruses are a ubiquitous family of nonpathogenic retroviruses that have potential as gene therapy vectors in humans. Here we demonstrate that bovine foamy virus (BFV) expresses high levels of three viral microRNAs (miRNAs) in BFV-infected cells in culture and also in infected cattle. The BFV miRNAs are unusual in that they are initially transcribed by RNA polymerase III as a single, ∼ 122-nt pri-miRNA that is subsequently processed to release three fully functional miRNAs. The observation that BFV, a foamy virus, is able to express viral miRNAs in infected cells adds to emerging evidence that miRNA expression is a common, albeit clearly not universal, property of retroviruses and suggests that these miRNAs may exert a significant effect on viral replication in vivo.
Figures

Similar articles
-
Molecular Epidemiology and Whole-Genome Analysis of Bovine Foamy Virus in Japan.Viruses. 2021 May 28;13(6):1017. doi: 10.3390/v13061017. Viruses. 2021. PMID: 34071542 Free PMC article.
-
Bovine Foamy Virus: Shared and Unique Molecular Features In Vitro and In Vivo.Viruses. 2019 Nov 21;11(12):1084. doi: 10.3390/v11121084. Viruses. 2019. PMID: 31766538 Free PMC article. Review.
-
Functional characterization of the bovine foamy virus miRNA expression cassette and its dumbbell-shaped pri-miRNA.Virus Genes. 2018 Aug;54(4):550-560. doi: 10.1007/s11262-018-1574-z. Epub 2018 May 31. Virus Genes. 2018. PMID: 29855776
-
Functional Analyses of Bovine Foamy Virus-Encoded miRNAs Reveal the Importance of a Defined miRNA for Virus Replication and Host-Virus Interaction.Viruses. 2020 Nov 2;12(11):1250. doi: 10.3390/v12111250. Viruses. 2020. PMID: 33147813 Free PMC article.
-
Non-primate foamy viruses.Curr Top Microbiol Immunol. 2003;277:197-211. doi: 10.1007/978-3-642-55701-9_9. Curr Top Microbiol Immunol. 2003. PMID: 12908774 Review.
Cited by
-
Bovine Leukemia Virus Small Noncoding RNAs Are Functional Elements That Regulate Replication and Contribute to Oncogenesis In Vivo.PLoS Pathog. 2016 Apr 28;12(4):e1005588. doi: 10.1371/journal.ppat.1005588. eCollection 2016 Apr. PLoS Pathog. 2016. PMID: 27123579 Free PMC article.
-
Identification of tri-phosphatase activity in the biogenesis of retroviral microRNAs and RNAP III-generated shRNAs.Nucleic Acids Res. 2014 Dec 16;42(22):13949-62. doi: 10.1093/nar/gku1247. Epub 2014 Nov 26. Nucleic Acids Res. 2014. PMID: 25428356 Free PMC article.
-
Interplay between SARS-CoV-2-derived miRNAs, immune system, vitamin D pathway and respiratory system.J Cell Mol Med. 2021 Aug;25(16):7825-7839. doi: 10.1111/jcmm.16694. Epub 2021 Jun 22. J Cell Mol Med. 2021. PMID: 34159729 Free PMC article.
-
Molecular Epidemiology and Whole-Genome Analysis of Bovine Foamy Virus in Japan.Viruses. 2021 May 28;13(6):1017. doi: 10.3390/v13061017. Viruses. 2021. PMID: 34071542 Free PMC article.
-
RNA Virus-Encoded miRNAs: Current Insights and Future Challenges.Front Microbiol. 2021 Jun 24;12:679210. doi: 10.3389/fmicb.2021.679210. eCollection 2021. Front Microbiol. 2021. PMID: 34248890 Free PMC article. Review.
References
-
- Parameswaran P, Sklan E, Wilkins C, Burgon T, Samuel MA, Lu R, Ansel KM, Heissmeyer V, Einav S, Jackson W, Doukas T, Paranjape S, Polacek C, dos Santos FB, Jalili R, Babrzadeh F, Gharizadeh B, Grimm D, Kay M, Koike S, Sarnow P, Ronaghi M, Ding SW, Harris E, Chow M, Diamond MS, Kirkegaard K, Glenn JS, Fire AZ. 2010. Six RNA viruses and forty-one hosts: viral small RNAs and modulation of small RNA repertoires in vertebrate and invertebrate systems. PLoS Pathog. 6:e1000764. 10.1371/journal.ppat.1000764 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

