The human platelet: strong transcriptome correlations among individuals associate weakly with the platelet proteome
- PMID: 24524654
- PMCID: PMC3937023
- DOI: 10.1186/1745-6150-9-3
The human platelet: strong transcriptome correlations among individuals associate weakly with the platelet proteome
Abstract
Background: For the anucleate platelet it has been unclear how well platelet transcriptomes correlate among different donors or across different RNA profiling platforms, and what the transcriptomes' relationship is with the platelet proteome. We profiled the platelet transcriptome of 10 healthy young males (5 white and 5 black) with no notable clinical history using RNA sequencing and by Affymetrix microarray.
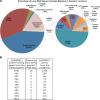
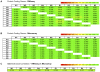
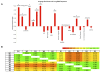
Results: We found that the abundance of platelet mRNA transcripts was highly correlated across the 10 individuals, independently of race and of the employed technology. Our RNA-seq data showed that these high inter-individual correlations extend beyond mRNAs to several categories of non-coding RNAs. Pseudogenes represented a notable exception by exhibiting a difference in expression by race. Comparison of our mRNA signatures to a publicly available quantitative platelet proteome showed that most (87.5%) identified platelet proteins had a detectable corresponding mRNA. However, a high number of mRNAs that were present in the transcriptomes of all 10 individuals had no representation in the proteome. Spearman correlations of the relative abundances for those genes represented by both an mRNA and a protein showed a weak (~0.3) connection. Further analysis of the overlapping and non-overlapping platelet mRNAs and proteins identified gene groups corresponding to distinct cellular processes.
Conclusions: The results of our analyses provide novel insights for platelet biology, show only a weak connection between the platelet transcriptome and proteome, and indicate that it is feasible to assemble a platelet mRNA-ome that can serve as a reference for future platelet transcriptomic studies of human health and disease.
Figures


References
-
- Cecchetti L, Tolley ND, Michetti N, Bury L, Weyrich AS, Gresele P. Megakaryocytes differentially sort mRNAs for matrix metalloproteinases and their inhibitors into platelets: a mechanism for regulating synthetic events. Blood. 2011;9:1903–1911. doi: 10.1182/blood-2010-12-324517. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

