A LC3-interacting motif in the influenza A virus M2 protein is required to subvert autophagy and maintain virion stability
- PMID: 24528869
- PMCID: PMC3991421
- DOI: 10.1016/j.chom.2014.01.006
A LC3-interacting motif in the influenza A virus M2 protein is required to subvert autophagy and maintain virion stability
Abstract
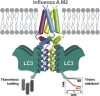
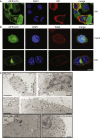
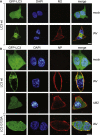
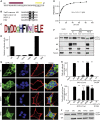
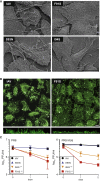
Autophagy recycles cellular components and defends cells against intracellular pathogens. While viruses must evade autophagocytic destruction, some viruses can also subvert autophagy for their own benefit. The ability of influenza A virus (IAV) to evade autophagy depends on the Matrix 2 (M2) ion-channel protein. We show that the cytoplasmic tail of IAV M2 interacts directly with the essential autophagy protein LC3 and promotes LC3 relocalization to the unexpected destination of the plasma membrane. LC3 binding is mediated by a highly conserved LC3-interacting region (LIR) in M2. The M2 LIR is required for LC3 redistribution to the plasma membrane in virus-infected cells. Mutations in M2 that abolish LC3 binding interfere with filamentous budding and reduce virion stability. IAV therefore subverts autophagy by mimicking a host short linear protein-protein interaction motif. This strategy may facilitate transmission of infection between organisms by enhancing the stability of viral progeny.
Copyright © 2014 Elsevier Inc. All rights reserved.
Figures
Comment in
-
Influenza A virus lures autophagic protein LC3 to budding sites.Cell Host Microbe. 2014 Feb 12;15(2):130-1. doi: 10.1016/j.chom.2014.01.014. Cell Host Microbe. 2014. PMID: 24528859
References
-
- Alemu E.A., Lamark T., Torgersen K.M., Birgisdottir A.B., Larsen K.B., Jain A., Olsvik H., Øvervatn A., Kirkin V., Johansen T. ATG8 family proteins act as scaffolds for assembly of the ULK complex: sequence requirements for LC3-interacting region (LIR) motifs. J. Biol. Chem. 2012;287:39275–39290. - PMC - PubMed
-
- Barrios-Rodiles M., Brown K.R., Ozdamar B., Bose R., Liu Z., Donovan R.S., Shinjo F., Liu Y., Dembowy J., Taylor I.W. High-throughput mapping of a dynamic signaling network in mammalian cells. Science. 2005;307:1621–1625. - PubMed
-
- Bourmakina S.V., García-Sastre A. Reverse genetics studies on the filamentous morphology of influenza A virus. J. Gen. Virol. 2003;84:517–527. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

