Role of F1C fimbriae, flagella, and secreted bacterial components in the inhibitory effect of probiotic Escherichia coli Nissle 1917 on atypical enteropathogenic E. coli infection
- PMID: 24549324
- PMCID: PMC3993448
- DOI: 10.1128/IAI.01431-13
Role of F1C fimbriae, flagella, and secreted bacterial components in the inhibitory effect of probiotic Escherichia coli Nissle 1917 on atypical enteropathogenic E. coli infection
Abstract
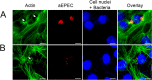
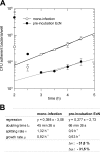
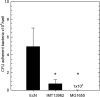
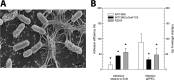
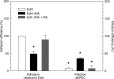
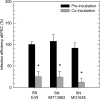
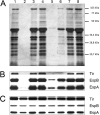
Enteropathogenic Escherichia coli (EPEC) is recognized as an important intestinal pathogen that frequently causes acute and persistent diarrhea in humans and animals. The use of probiotic bacteria to prevent diarrhea is gaining increasing interest. The probiotic E. coli strain Nissle 1917 (EcN) is known to be effective in the treatment of several gastrointestinal disorders. While both in vitro and in vivo studies have described strong inhibitory effects of EcN on enteropathogenic bacteria, including pathogenic E. coli, the underlying molecular mechanisms remain largely unknown. In this study, we examined the inhibitory effect of EcN on infections of porcine intestinal epithelial cells with atypical enteropathogenic E. coli (aEPEC) with respect to single infection steps, including adhesion, microcolony formation, and the attaching and effacing phenotype. We show that EcN drastically reduced the infection efficiencies of aEPEC by inhibiting bacterial adhesion and growth of microcolonies, but not the attaching and effacing of adherent bacteria. The inhibitory effect correlated with EcN adhesion capacities and was predominantly mediated by F1C fimbriae, but also by H1 flagella, which served as bridges between EcN cells. Furthermore, EcN seemed to interfere with the initial adhesion of aEPEC to host cells by secretion of inhibitory components. These components do not appear to be specific to EcN, but we propose that the strong adhesion capacities enable EcN to secrete sufficient local concentrations of the inhibitory factors. The results of this study are consistent with a mode of action whereby EcN inhibits secretion of virulence-associated proteins of EPEC, but not their expression.
Figures



Similar articles
-
E. coli Nissle 1917 Affects Salmonella adhesion to porcine intestinal epithelial cells.PLoS One. 2011 Feb 17;6(2):e14712. doi: 10.1371/journal.pone.0014712. PLoS One. 2011. PMID: 21379575 Free PMC article.
-
Interleukin-8, CXCL1, and MicroRNA miR-146a Responses to Probiotic Escherichia coli Nissle 1917 and Enteropathogenic E. coli in Human Intestinal Epithelial T84 and Monocytic THP-1 Cells after Apical or Basolateral Infection.Infect Immun. 2016 Aug 19;84(9):2482-92. doi: 10.1128/IAI.00402-16. Print 2016 Sep. Infect Immun. 2016. PMID: 27297392 Free PMC article.
-
Porcine E. coli: virulence-associated genes, resistance genes and adhesion and probiotic activity tested by a new screening method.PLoS One. 2013 Apr 26;8(4):e59242. doi: 10.1371/journal.pone.0059242. Print 2013. PLoS One. 2013. PMID: 23658605 Free PMC article.
-
[Molecular pathogenesis, epidemiology and diagnosis of enteropathogenic Escherichia coli].Salud Publica Mex. 2007 Sep-Oct;49(5):376-86. doi: 10.1590/s0036-36342007000500008. Salud Publica Mex. 2007. PMID: 17952245 Review. Spanish.
-
Enteropathogenic Escherichia coli: foe or innocent bystander?Clin Microbiol Infect. 2015 Aug;21(8):729-34. doi: 10.1016/j.cmi.2015.01.015. Epub 2015 Jan 28. Clin Microbiol Infect. 2015. PMID: 25726041 Free PMC article. Review.
Cited by
-
Role of Aeromonas hydrophila flagella glycosylation in adhesion to Hep-2 cells, biofilm formation and immune stimulation.Int J Mol Sci. 2014 Nov 28;15(12):21935-46. doi: 10.3390/ijms151221935. Int J Mol Sci. 2014. PMID: 25464381 Free PMC article.
-
A Newly Isolated Bacillus subtilis Strain Named WS-1 Inhibited Diarrhea and Death Caused by Pathogenic Escherichia coli in Newborn Piglets.Front Microbiol. 2019 Jun 12;10:1248. doi: 10.3389/fmicb.2019.01248. eCollection 2019. Front Microbiol. 2019. PMID: 31249559 Free PMC article.
-
Escherichia coli ST302: Genomic Analysis of Virulence Potential and Antimicrobial Resistance Mediated by Mobile Genetic Elements.Front Microbiol. 2020 Jan 21;10:3098. doi: 10.3389/fmicb.2019.03098. eCollection 2019. Front Microbiol. 2020. PMID: 32063891 Free PMC article.
-
Lactobacillus johnsonii L531 Ameliorates Escherichia coli-Induced Cell Damage via Inhibiting NLRP3 Inflammasome Activity and Promoting ATG5/ATG16L1-Mediated Autophagy in Porcine Mammary Epithelial Cells.Vet Sci. 2020 Aug 14;7(3):112. doi: 10.3390/vetsci7030112. Vet Sci. 2020. PMID: 32823867 Free PMC article.
-
Effects of an orally supplemented probiotic on the autophagy protein LC3 and Beclin1 in placentas undergoing spontaneous delivery during normal pregnancy.BMC Pregnancy Childbirth. 2020 Apr 15;20(1):216. doi: 10.1186/s12884-020-02905-z. BMC Pregnancy Childbirth. 2020. PMID: 32295534 Free PMC article.
References
-
- Bertschinger HU, Fairbrother JM. 1999. Escherichia coli infections, p 431–468 In Straw BE, D'Allaire S, Mengeling WL, Taylor DJ. (ed), Diseases of swine, 8th ed. Blackwell Science, Oxford, United Kingdom
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

