IcgA is a virulence factor of Rhodococcus equi that modulates intracellular growth
- PMID: 24549327
- PMCID: PMC3993432
- DOI: 10.1128/IAI.01670-13
IcgA is a virulence factor of Rhodococcus equi that modulates intracellular growth
Abstract
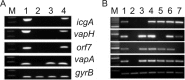
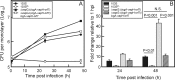
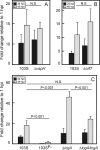
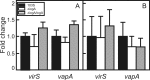
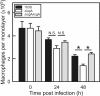
Virulence of the intracellular pathogen Rhodococcus equi depends on a 21.3-kb pathogenicity island located on a conjugative plasmid. To date, the only nonregulatory pathogenicity island-encoded virulence factor identified is the cell envelope-associated VapA protein. Although the pathogenicity islands from porcine and equine R. equi isolates have undergone major rearrangements, the virR operon (virR-icgA-vapH-orf7-virS) is highly conserved in both, suggesting these genes play an important role in pathogenicity. VirR and VirS are transcriptional regulators controlling expression of pathogenicity island genes, including vapA. Here, we show that while vapH and orf7 are dispensable for intracellular growth of R. equi, deletion of icgA, formerly known as orf5, encoding a major facilitator superfamily transport protein, elicited an enhanced growth phenotype in macrophages and a significant reduction in macrophage viability, while extracellular growth in broth remained unaffected. Transcription of virS, located downstream of icgA, and vapA was not affected by the icgA deletion during growth in broth or in macrophages, showing that the enhanced growth phenotype caused by deletion of icgA was not mediated through abnormal transcription of these genes. Transcription of icgA increased 6-fold within 2 h following infection of macrophages and remained significantly higher 48 h postinfection compared to levels at the start of the infection. The major facilitator superfamily transport protein IcgA is the first factor identified in R. equi that negatively affects intracellular replication. Aside from VapA, it is only the second pathogenicity island-encoded structural protein shown to play a direct role in intracellular growth of this pathogenic actinomycete.
Figures


Similar articles
-
Transcriptome reprogramming by plasmid-encoded transcriptional regulators is required for host niche adaption of a macrophage pathogen.Infect Immun. 2015 Aug;83(8):3137-45. doi: 10.1128/IAI.00230-15. Epub 2015 May 26. Infect Immun. 2015. PMID: 26015480 Free PMC article.
-
Characterization of the role of the pathogenicity island and vapG in the virulence of the intracellular actinomycete pathogen Rhodococcus equi.Infect Immun. 2010 Aug;78(8):3323-34. doi: 10.1128/IAI.00081-10. Epub 2010 May 3. Infect Immun. 2010. PMID: 20439471 Free PMC article.
-
The vapA co-expressed virulence plasmid gene vcgB (orf10) of the intracellular actinomycete Rhodococcus equi.Microbiology (Reading). 2011 Aug;157(Pt 8):2357-2368. doi: 10.1099/mic.0.049759-0. Epub 2011 May 12. Microbiology (Reading). 2011. PMID: 21565932
-
Pathogenesis and virulence of Rhodococcus equi.Vet Microbiol. 1997 Jun 16;56(3-4):257-68. doi: 10.1016/s0378-1135(97)00094-1. Vet Microbiol. 1997. PMID: 9226840 Review.
-
Molecular and infection biology of the horse pathogen Rhodococcus equi.FEMS Microbiol Rev. 2009 Sep;33(5):870-91. doi: 10.1111/j.1574-6976.2009.00181.x. Epub 2009 Apr 23. FEMS Microbiol Rev. 2009. PMID: 19453748 Review.
Cited by
-
Transcriptome reprogramming by plasmid-encoded transcriptional regulators is required for host niche adaption of a macrophage pathogen.Infect Immun. 2015 Aug;83(8):3137-45. doi: 10.1128/IAI.00230-15. Epub 2015 May 26. Infect Immun. 2015. PMID: 26015480 Free PMC article.
-
The extracellular thioredoxin Etrx3 is required for macrophage infection in Rhodococcus equi.Vet Res. 2020 Mar 10;51(1):38. doi: 10.1186/s13567-020-00763-3. Vet Res. 2020. PMID: 32156317 Free PMC article.
-
The N-terminal domain is required for cell surface localisation of VapA, a member of the Vap family of Rhodococcus equi virulence proteins.PLoS One. 2024 Feb 29;19(2):e0298900. doi: 10.1371/journal.pone.0298900. eCollection 2024. PLoS One. 2024. PMID: 38421980 Free PMC article.
-
LysR-Type Transcriptional Regulator VirR Responds to Temperature and pH and Directly Activates the Transcription of virS-Containing Operon in Rhodococcus equi.Int J Microbiol. 2025 Jan 3;2025:6618952. doi: 10.1155/ijm/6618952. eCollection 2025. Int J Microbiol. 2025. PMID: 39802684 Free PMC article.
-
The pathogenic actinobacterium Rhodococcus equi: what's in a name?Mol Microbiol. 2019 Jul;112(1):1-15. doi: 10.1111/mmi.14267. Epub 2019 Jun 17. Mol Microbiol. 2019. PMID: 31099908 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

