Trithorax monomethylates histone H3K4 and interacts directly with CBP to promote H3K27 acetylation and antagonize Polycomb silencing
- PMID: 24550119
- PMCID: PMC3929413
- DOI: 10.1242/dev.102392
Trithorax monomethylates histone H3K4 and interacts directly with CBP to promote H3K27 acetylation and antagonize Polycomb silencing
Abstract
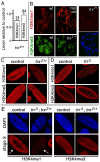
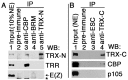
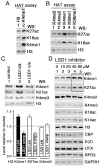
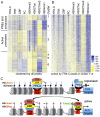
Trithorax (TRX) antagonizes epigenetic silencing by Polycomb group (PcG) proteins, stimulates enhancer-dependent transcription, and establishes a 'cellular memory' of active transcription of PcG-regulated genes. The mechanisms underlying these TRX functions remain largely unknown, but are presumed to involve its histone H3K4 methyltransferase activity. We report that the SET domains of TRX and TRX-related (TRR) have robust histone H3K4 monomethyltransferase activity in vitro and that Tyr3701 of TRX and Tyr2404 of TRR prevent them from being trimethyltransferases. The trx(Z11) missense mutation (G3601S), which abolishes H3K4 methyltransferase activity in vitro, reduces the H3K4me1 but not the H3K4me3 level in vivo. trx(Z11) also suppresses the impaired silencing phenotypes of the Pc(3) mutant, suggesting that H3K4me1 is involved in antagonizing Polycomb silencing. Polycomb silencing is also antagonized by TRX-dependent H3K27 acetylation by CREB-binding protein (CBP). We show that perturbation of Polycomb silencing by TRX overexpression requires CBP. We also show that TRX and TRR are each physically associated with CBP in vivo, that TRX binds directly to the CBP KIX domain, and that the chromatin binding patterns of TRX and TRR are highly correlated with CBP and H3K4me1 genome-wide. In vitro acetylation of H3K27 by CBP is enhanced on K4me1-containing H3 substrates, and independently altering the H3K4me1 level in vivo, via the H3K4 demethylase LSD1, produces concordant changes in H3K27ac. These data indicate that the catalytic activities of TRX and CBP are physically coupled and suggest that both activities play roles in antagonizing Polycomb silencing, stimulating enhancer activity and cellular memory.
Keywords: CBP; Drosophila; H3K4 monomethylation; Trithorax.
Figures




References
-
- Arnold C. D., Gerlach D., Stelzer C., Boryń L. M., Rath M., Stark A. (2013). Genome-wide quantitative enhancer activity maps identified by STARR-seq. Science 339, 1074–1077 - PubMed
-
- Bonn S., Zinzen R. P., Girardot C., Gustafson E. H., Perez-Gonzalez A., Delhomme N., Ghavi-Helm Y., Wilczyński B., Riddell A., Furlong E. E. (2012). Tissue-specific analysis of chromatin state identifies temporal signatures of enhancer activity during embryonic development. Nat. Genet. 44, 148–156 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

