The hippocampal CA2 ensemble is sensitive to contextual change
- PMID: 24553945
- PMCID: PMC6608518
- DOI: 10.1523/JNEUROSCI.2563-13.2014
The hippocampal CA2 ensemble is sensitive to contextual change
Abstract
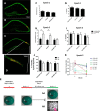
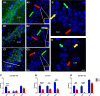
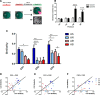
Contextual learning involves associating cues with an environment and relating them to past experience. Previous data indicate functional specialization within the hippocampal circuit: the dentate gyrus (DG) is crucial for discriminating similar contexts, whereas CA3 is required for associative encoding and recall. Here, we used Arc/H1a catFISH imaging to address the contribution of the largely overlooked CA2 region to contextual learning by comparing ensemble codes across CA3, CA2, and CA1 in mice exposed to familiar, altered, and novel contexts. Further, to manipulate the quality of information arriving in CA2 we used two hippocampal mutant mouse lines, CA3-NR1 KOs and DG-NR1 KOs, that result in hippocampal CA3 neuronal activity that is uncoupled from the animal's sensory environment. Our data reveal largely coherent responses across the CA axis in control mice in purely novel or familiar contexts; however, in the mutant mice subject to these protocols the CA2 response becomes uncoupled from CA1 and CA3. Moreover, we show in wild-type mice that the CA2 ensemble is more sensitive than CA1 and CA3 to small changes in overall context. Our data suggest that CA2 may be tuned to remap in response to any conflict between stored and current experience.
Keywords: Arc; CA2; H1a; IEG; context learning; hippocampus.
Figures

References
-
- Carninci P, Carninci P, Kasukawa T, Katayama S, Gough J, Frith MC, Maeda N, Oyama R, Ravasi T, Lenhard B, Wells C, Kodzius R, Shimokawa K, Bajic VB, Brenner SE, Batalov S, Forrest AR, Zavolan M, Davis MJ, Wilming LG, et al. The transcriptional landscape of the mammalian genome. Science. 2005;309:1559–1563. doi: 10.1126/science.1112014. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous
