Genome-wide binding patterns of thyroid hormone receptor beta
- PMID: 24558356
- PMCID: PMC3928038
- DOI: 10.1371/journal.pone.0081186
Genome-wide binding patterns of thyroid hormone receptor beta
Abstract
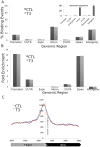
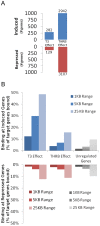
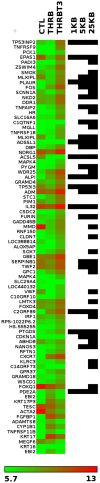
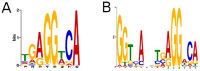
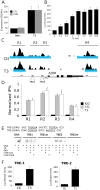
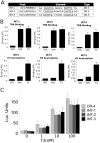
Thyroid hormone (TH) receptors (TRs) play central roles in metabolism and are major targets for pharmaceutical intervention. Presently, however, there is limited information about genome wide localizations of TR binding sites. Thus, complexities of TR genomic distribution and links between TRβ binding events and gene regulation are not fully appreciated. Here, we employ a BioChIP approach to capture TR genome-wide binding events in a liver cell line (HepG2). Like other NRs, TRβ appears widely distributed throughout the genome. Nevertheless, there is striking enrichment of TRβ binding sites immediately 5' and 3' of transcribed genes and TRβ can be detected near 50% of T3 induced genes. In contrast, no significant enrichment of TRβ is seen at negatively regulated genes or genes that respond to unliganded TRs in this system. Canonical TRE half-sites are present in more than 90% of TRβ peaks and classical TREs are also greatly enriched, but individual TRE organization appears highly variable with diverse half-site orientation and spacing. There is also significant enrichment of binding sites for TR associated transcription factors, including AP-1 and CTCF, near TR peaks. We conclude that T3-dependent gene induction commonly involves proximal TRβ binding events but that far-distant binding events are needed for T3 induction of some genes and that distinct, indirect, mechanisms are often at play in negative regulation and unliganded TR actions. Better understanding of genomic context of TR binding sites will help us determine why TR regulates genes in different ways and determine possibilities for selective modulation of TR action.
Conflict of interest statement
Figures

References
-
- Nielsen R, Pedersen TA, Hagenbeek D, Moulos P, Siersbaek R, et al. (2008) Genome-wide profiling of PPARgamma:RXR and RNA polymerase II occupancy reveals temporal activation of distinct metabolic pathways and changes in RXR dimer composition during adipogenesis. Genes & development 22 (21) 2953–67. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

