Deciphering the rules by which dynamics of mRNA secondary structure affect translation efficiency in Saccharomyces cerevisiae
- PMID: 24561808
- PMCID: PMC4005662
- DOI: 10.1093/nar/gku159
Deciphering the rules by which dynamics of mRNA secondary structure affect translation efficiency in Saccharomyces cerevisiae
Abstract
Messenger RNA (mRNA) secondary structure decreases the elongation rate, as ribosomes must unwind every structure they encounter during translation. Therefore, the strength of mRNA secondary structure is assumed to be reduced in highly translated mRNAs. However, previous studies in vitro reported a positive correlation between mRNA folding strength and protein abundance. The counterintuitive finding suggests that mRNA secondary structure affects translation efficiency in an undetermined manner. Here, we analyzed the folding behavior of mRNA during translation and its effect on translation efficiency. We simulated translation process based on a novel computational model, taking into account the interactions among ribosomes, codon usage and mRNA secondary structures. We showed that mRNA secondary structure shortens ribosomal distance through the dynamics of folding strength. Notably, when adjacent ribosomes are close, mRNA secondary structures between them disappear, and codon usage determines the elongation rate. More importantly, our results showed that the combined effect of mRNA secondary structure and codon usage in highly translated mRNAs causes a short ribosomal distance in structural regions, which in turn eliminates the structures during translation, leading to a high elongation rate. Together, these findings reveal how the dynamics of mRNA secondary structure coupling with codon usage affect translation efficiency.
Figures
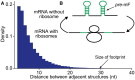
 , and release proteins with rate
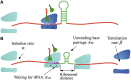
, and release proteins with rate  . During elongation, translating ribosomes wait for their cognate tRNAs at position i and simultaneously unwind the base pairings located at position i + L/2 (L = 42 nt). (A) shows the structure that the first ribosome encounters (the first ribosome is blue). (B) shows the structure that the third ribosome encounters. mRNA secondary structure is weakened due to the constraints of ribosomes. In our model, we assume that different ribosomes might encounter the structure with different folding strength at the same site because the pattern of mRNA folding might be changed when ribosomes have bound to mRNA (see text for details).
. During elongation, translating ribosomes wait for their cognate tRNAs at position i and simultaneously unwind the base pairings located at position i + L/2 (L = 42 nt). (A) shows the structure that the first ribosome encounters (the first ribosome is blue). (B) shows the structure that the third ribosome encounters. mRNA secondary structure is weakened due to the constraints of ribosomes. In our model, we assume that different ribosomes might encounter the structure with different folding strength at the same site because the pattern of mRNA folding might be changed when ribosomes have bound to mRNA (see text for details).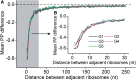
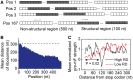
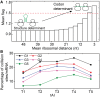
 (see ‘Materials and Methods’ section for the calculation of
(see ‘Materials and Methods’ section for the calculation of  ) among the five groups when ribosomal distance is longer than 5 nt. (B) When the distance is shorter than 5 nt, there is no structure between adjacent ribosomes,
) among the five groups when ribosomal distance is longer than 5 nt. (B) When the distance is shorter than 5 nt, there is no structure between adjacent ribosomes,  is determined by pre-mF strength.
is determined by pre-mF strength.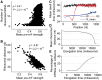
Similar articles
-
Translation elongation and mRNA stability are coupled through the ribosomal A-site.RNA. 2018 Oct;24(10):1377-1389. doi: 10.1261/rna.066787.118. Epub 2018 Jul 11. RNA. 2018. PMID: 29997263 Free PMC article.
-
Translation efficiency is determined by both codon bias and folding energy.Proc Natl Acad Sci U S A. 2010 Feb 23;107(8):3645-50. doi: 10.1073/pnas.0909910107. Epub 2010 Feb 2. Proc Natl Acad Sci U S A. 2010. PMID: 20133581 Free PMC article.
-
Codon Usage Influences the Local Rate of Translation Elongation to Regulate Co-translational Protein Folding.Mol Cell. 2015 Sep 3;59(5):744-54. doi: 10.1016/j.molcel.2015.07.018. Epub 2015 Aug 27. Mol Cell. 2015. PMID: 26321254 Free PMC article.
-
Synonymous but Not Silent: The Codon Usage Code for Gene Expression and Protein Folding.Annu Rev Biochem. 2021 Jun 20;90:375-401. doi: 10.1146/annurev-biochem-071320-112701. Epub 2021 Jan 13. Annu Rev Biochem. 2021. PMID: 33441035 Free PMC article.
-
When mRNA translation meets decay.Biochem Soc Trans. 2017 Apr 15;45(2):339-351. doi: 10.1042/BST20160243. Biochem Soc Trans. 2017. PMID: 28408474 Review.
Cited by
-
RPLP1 and RPLP2 Are Essential Flavivirus Host Factors That Promote Early Viral Protein Accumulation.J Virol. 2017 Jan 31;91(4):e01706-16. doi: 10.1128/JVI.01706-16. Print 2017 Feb 15. J Virol. 2017. PMID: 27974556 Free PMC article.
-
Utilizing 5' UTR Engineering Enables Fine-Tuning of Multiple Genes within Operons to Balance Metabolic Flux in Bacillus subtilis.Biology (Basel). 2024 Apr 19;13(4):277. doi: 10.3390/biology13040277. Biology (Basel). 2024. PMID: 38666889 Free PMC article.
-
Readthrough-induced misincorporated amino acid ratios guide mutant-specific therapeutic approaches for two CFTR nonsense mutations.Front Pharmacol. 2024 Apr 25;15:1389586. doi: 10.3389/fphar.2024.1389586. eCollection 2024. Front Pharmacol. 2024. PMID: 38725656 Free PMC article.
-
RNA search engines empower the bacterial intranet.Biochem Soc Trans. 2017 Aug 15;45(4):987-997. doi: 10.1042/BST20160373. Epub 2017 Jul 14. Biochem Soc Trans. 2017. PMID: 28710287 Free PMC article. Review.
-
GCN sensitive protein translation in yeast.PLoS One. 2020 Sep 18;15(9):e0233197. doi: 10.1371/journal.pone.0233197. eCollection 2020. PLoS One. 2020. PMID: 32946445 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

