A plastid without a genome: evidence from the nonphotosynthetic green algal genus Polytomella
- PMID: 24563281
- PMCID: PMC3982744
- DOI: 10.1104/pp.113.233718
A plastid without a genome: evidence from the nonphotosynthetic green algal genus Polytomella
Abstract
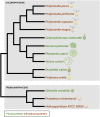
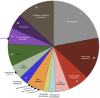
Polytomella spp. are free-living, nonphotosynthetic green algae closely related to the model organism Chlamydomonas reinhardtii. Although colorless, Polytomella spp. have a plastid, but it is still unknown whether they harbor a plastid genome. We took a next generation sequencing approach, along with transcriptome sequencing, to search for a plastid genome and an associated gene expression system in Polytomella spp. Illumina sequencing of total DNA from four Polytomella spp. did not produce any recognizable plastid-derived reads but did generate a large number of mitochondrial DNA sequences. Transcriptomic analysis of Polytomella parva uncovered hundreds of putative nuclear-encoded, plastid-targeted proteins, which support the presence of plastid-based metabolic functions, similar to those observed in the plastids of other nonphotosynthetic algae. Conspicuously absent, however, were any plastid-targeted proteins involved in the expression, replication, or repair of plastid DNA. Based on these findings and earlier findings, we argue that the Polytomella genus represents the first well-supported example, to our knowledge, of a primary plastid-bearing lineage without a plastid genome.
Figures

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

