Phenotypic diversification by gene silencing in Phytophthora plant pathogens
- PMID: 24563702
- PMCID: PMC3917941
- DOI: 10.4161/cib.25890
Phenotypic diversification by gene silencing in Phytophthora plant pathogens
Abstract
Advances in genome sequencing technologies have enabled generation of unprecedented information on genome content and organization. Eukaryote genomes in particular may contain large populations of transposable elements (TEs) and other repeated sequences. Active TEs can result in insertional mutations, altered transcription levels and ectopic recombination of DNA. The genome of the oomycete plant pathogen, Phytophthora infestans, contains vast numbers of TE sequences. There are also hundreds of predicted disease-promoting effector proteins, predominantly located in TE-rich genomic regions. Expansion of effector gene families is also a genomic signature of related oomycetes such as P. sojae. Deep sequencing of small RNAs (sRNAs) from P. infestans has identified sRNAs derived from all families of transposons, highlighting the importance of RNA silencing for maintaining these genomic invaders in an inactive form. Small RNAs were also identified from specific effector encoding genes, possibly leading to RNA silencing of these genes and variation in pathogenicity and virulence toward plant resistance genes. Similar findings have also recently been made for the distantly related species, P. sojae. Small RNA "hotspots" originating from arrays of amplified gene sequences, or from genes displaying overlapping antisense transcription, were also identified in P. infestans. These findings suggest a major role for RNA silencing processes in the adaptability and diversification of these economically important plant pathogens. Here we review the latest progress and understanding of gene silencing in oomycetes with emphasis on transposable elements and sRNA-associated events.
Keywords: Phytophthora; effectors; oomycete; small RNAs; transposable element.
Figures
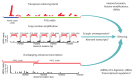
References
-
- Raffaele S, Kamoun S. Genome evolution in filamentous plant pathogens: why bigger can be better. Nat Rev Microbiol. 2012;10:417–30. - PubMed
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources
