A histidine aspartate ionic lock gates the iron passage in miniferritins from Mycobacterium smegmatis
- PMID: 24573673
- PMCID: PMC4036245
- DOI: 10.1074/jbc.M113.524421
A histidine aspartate ionic lock gates the iron passage in miniferritins from Mycobacterium smegmatis
Abstract
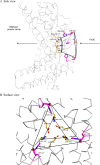
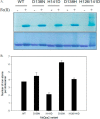
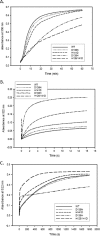
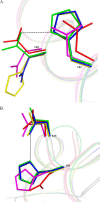
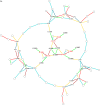
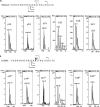
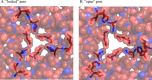
Dps (DNA-binding protein from starved cells) are dodecameric assemblies belonging to the ferritin family that can bind DNA, carry out ferroxidation, and store iron in their shells. The ferritin-like trimeric pore harbors the channel for the entry and exit of iron. By representing the structure of Dps as a network we have identified a charge-driven interface formed by a histidine aspartate cluster at the pore interface unique to Mycobacterium smegmatis Dps protein, MsDps2. Site-directed mutagenesis was employed to generate mutants to disrupt the charged interactions. Kinetics of iron uptake/release of the wild type and mutants were compared. Crystal structures were solved at a resolution of 1.8-2.2 Å for the various mutants to compare structural alterations vis à vis the wild type protein. The substitutions at the pore interface resulted in alterations in the side chain conformations leading to an overall weakening of the interface network, especially in cases of substitutions that alter the charge at the pore interface. Contrary to earlier findings where conserved aspartate residues were found crucial for iron release, we propose here that in the case of MsDps2, it is the interplay of negative-positive potentials at the pore that enables proper functioning of the protein. In similar studies in ferritins, negative and positive patches near the iron exit pore were found to be important in iron uptake/release kinetics. The unique ionic cluster in MsDps2 makes it a suitable candidate to act as nano-delivery vehicle, as these gated pores can be manipulated to exhibit conformations allowing for slow or fast rates of iron release.
Keywords: DNA-binding Protein; Ion Channels; Ionic Cluster; Iron; Iron Oxidation; Mycobacterium; Mycobacterium smegmatis; X-ray Crystallography.
Figures




References
-
- Aisen P., Enns C., Wessling-Resnick M. (2001) Chemistry and biology of eukaryotic iron metabolism. Int. J. Biochem. Cell Biol. 33, 940–959 - PubMed
-
- Benjamin J. A., Desnoyers G., Morissette A., Salvail H., Massé E. (2010) Dealing with oxidative stress and iron starvation in microorganisms: an overview. Can. J. Physiol. Pharmacol. 88, 264–272 - PubMed
-
- Bozzi M., Mignogna G., Stefanini S., Barra D., Longhi C., Valenti P., Chiancone E. (1997) A novel non-heme iron-binding ferritin related to the DNA-binding proteins of the Dps family in Listeria innocua. J. Biol. Chem. 272, 3259–3265 - PubMed
-
- Zhao G., Ceci P., Ilari A., Giangiacomo L., Laue T. M., Chiancone E., Chasteen N. D. (2002) Iron and hydrogen peroxide detoxification properties of DNA-binding protein from starved cells. A ferritin-like DNA-binding protein of Escherichia coli. J. Biol. Chem. 277, 27689–27696 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

