Co-culture with periodontal ligament stem cells enhances osteogenic gene expression in de-differentiated fat cells
- PMID: 24573839
- PMCID: PMC4186972
- DOI: 10.1007/s13577-014-0091-1
Co-culture with periodontal ligament stem cells enhances osteogenic gene expression in de-differentiated fat cells
Abstract
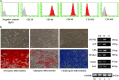
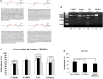
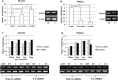
In recent decades, de-differentiated fat cells (DFAT cells) have emerged in regenerative medicine because of their trans-differentiation capability and the fact that their characteristics are similar to bone marrow mesenchymal stem cells. Even so, there is no evidence to support the osteogenic induction using DFAT cells in periodontal regeneration and also the co-culture system. Consequently, this study sought to evaluate the DFAT cells co-culture with periodontal ligament stem cells (PDLSCs) in vitro in terms of gene expression by comparing runt-related transcription factor 2 (RUNX2) and Peroxisome proliferator-activated receptor gamma 2 (PPARγ2) genes. We isolated DFAT cells from mature adipocytes and compared proliferation with PDLSCs. After co-culture with PDLSCs, we analyzed transcriptional activity implying by DNA methylation in all adipogenic gene promoters using combined bisulfite restriction analysis. We compared gene expression in RUNX2 gene with the PPARγ2 gene using quantitative RT-PCR. After being sub-cultured, DFAT cells demonstrated morphology similar to fibroblast-like cells. At the same time, PDLSCs established all stem cell characteristics. Interestingly, the co-culture system attenuated proliferation while enhancing osteogenic gene expression in RUNX2 gene. Using the co-culture system, DFAT cells could trans-differentiate into osteogenic lineage enhancing, but conversely, their adipogenic characteristic diminished. Therefore, DFAT cells and the co-culture system might be a novel cell-based therapy for promoting osteogenic differentiation in periodontal regeneration.
Figures


Similar articles
-
Effects of vascular endothelial cells on osteogenic differentiation of noncontact co-cultured periodontal ligament stem cells under hypoxia.J Periodontal Res. 2013 Feb;48(1):52-65. doi: 10.1111/j.1600-0765.2012.01503.x. Epub 2012 Aug 21. J Periodontal Res. 2013. PMID: 22905750
-
Canonical Wnt signaling differently modulates osteogenic differentiation of mesenchymal stem cells derived from bone marrow and from periodontal ligament under inflammatory conditions.Biochim Biophys Acta. 2014 Mar;1840(3):1125-34. doi: 10.1016/j.bbagen.2013.11.003. Epub 2013 Nov 12. Biochim Biophys Acta. 2014. PMID: 24231680
-
Downregulation of lncRNA DANCR promotes osteogenic differentiation of periodontal ligament stem cells.BMC Dev Biol. 2020 Jan 14;20(1):2. doi: 10.1186/s12861-019-0206-8. BMC Dev Biol. 2020. PMID: 31931700 Free PMC article.
-
Epigenetic regulation of osteogenic differentiation of periodontal ligament stem cells in periodontitis.Oral Dis. 2023 Oct;29(7):2529-2537. doi: 10.1111/odi.14491. Epub 2023 Jan 13. Oral Dis. 2023. PMID: 36582112 Review.
-
Insights into the molecular changes of adipocyte dedifferentiation and its future research opportunities.J Lipid Res. 2024 Nov;65(11):100644. doi: 10.1016/j.jlr.2024.100644. Epub 2024 Sep 18. J Lipid Res. 2024. PMID: 39303983 Free PMC article. Review.
Cited by
-
Epigenetic Regulation of Methylation in Determining the Fate of Dental Mesenchymal Stem Cells.Stem Cells Int. 2022 Sep 22;2022:5015856. doi: 10.1155/2022/5015856. eCollection 2022. Stem Cells Int. 2022. PMID: 36187229 Free PMC article. Review.
-
The Genetic Aspects of Periodontitis Pathogenesis and the Regenerative Properties of Stem Cells.Cells. 2024 Jan 9;13(2):117. doi: 10.3390/cells13020117. Cells. 2024. PMID: 38247810 Free PMC article. Review.
-
The Potential of Different Origin Stem Cells in Modulating Oral Bone Regeneration Processes.Cells. 2019 Jan 8;8(1):29. doi: 10.3390/cells8010029. Cells. 2019. PMID: 30625993 Free PMC article. Review.
-
Bone biomaterials and interactions with stem cells.Bone Res. 2017 Dec 21;5:17059. doi: 10.1038/boneres.2017.59. eCollection 2017. Bone Res. 2017. PMID: 29285402 Free PMC article. Review.
-
Long noncoding RNA HCG18 inhibits the differentiation of human bone marrow-derived mesenchymal stem cells in osteoporosis by targeting miR-30a-5p/NOTCH1 axis.Mol Med. 2020 Nov 11;26(1):106. doi: 10.1186/s10020-020-00219-6. Mol Med. 2020. PMID: 33176682 Free PMC article.
References
-
- Bartold PM, Xiao Y, Lyngstaadas SP, Paine ML, Snead ML. Principles and applications of cell delivery systems for periodontal regeneration. Periodontology. 2000;2006(41):123–135. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

