Characterization of epistatic interaction of QTLs LH8 and EH3 controlling heading date in rice
- PMID: 24584028
- PMCID: PMC3939452
- DOI: 10.1038/srep04263
Characterization of epistatic interaction of QTLs LH8 and EH3 controlling heading date in rice
Abstract
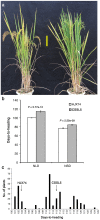
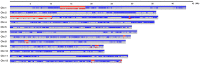
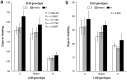
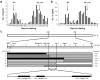
Heading date is a critical trait for adaptation of rice to different cultivation areas and cropping seasons. We evaluated the heading dates of 1,123 chromosome segments substitution lines (CSSLs) in the genetic background of an elite rice variety Huajingxian74 (HJX74). A CSSL with the substituted segments from Zihui100 exhibited late heading under both natural long-day (NLD) and natural short-day (NSD) conditions, and the late heading phenotype was controlled by two novel epistatic loci on chromosome 8 and chromosome 3, respectively, termed LH8 and EH3. The function of EH3 was dependent on the LH8 genotype through epistatic interaction between EH3(Zihui100) and LH8(Zihui100) alleles. Genetic and molecular characterization revealed LH8 encodes a CCAAT-box-binding transcription factor with Heading date1 (Hd1)-binding activity and may delay flowering by repressing the expression of Early heading date1 (Ehd1). Our work provides a solid foundation for further study on gene interaction in heading date and has application in breeding rice with greater adaptability.
Figures


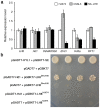

Similar articles
-
Hd16, a gene for casein kinase I, is involved in the control of rice flowering time by modulating the day-length response.Plant J. 2013 Oct;76(1):36-46. doi: 10.1111/tpj.12268. Epub 2013 Jul 25. Plant J. 2013. PMID: 23789941 Free PMC article.
-
DTH8 suppresses flowering in rice, influencing plant height and yield potential simultaneously.Plant Physiol. 2010 Aug;153(4):1747-58. doi: 10.1104/pp.110.156943. Epub 2010 Jun 21. Plant Physiol. 2010. PMID: 20566706 Free PMC article.
-
Analysis of Epistasis among QTLs on Heading Date based on Single Segment Substitution Lines in Rice.Sci Rep. 2018 Feb 15;8(1):3059. doi: 10.1038/s41598-018-20690-w. Sci Rep. 2018. PMID: 29449579 Free PMC article.
-
Transcriptional and post-transcriptional regulation of heading date in rice.New Phytol. 2021 May;230(3):943-956. doi: 10.1111/nph.17158. Epub 2021 Feb 14. New Phytol. 2021. PMID: 33341945 Free PMC article. Review.
-
Genome-wide association study of heading and flowering dates and construction of its prediction equation in Chinese common wheat.Theor Appl Genet. 2018 Nov;131(11):2271-2285. doi: 10.1007/s00122-018-3181-8. Epub 2018 Sep 14. Theor Appl Genet. 2018. PMID: 30218294 Review.
Cited by
-
Genetic control of flowering time in rice: integration of Mendelian genetics and genomics.Theor Appl Genet. 2016 Dec;129(12):2241-2252. doi: 10.1007/s00122-016-2773-4. Epub 2016 Sep 30. Theor Appl Genet. 2016. PMID: 27695876 Review.
-
Unconditional and conditional analysis of epistasis between tillering QTLs based on single segment substitution lines in rice.Sci Rep. 2020 Sep 28;10(1):15912. doi: 10.1038/s41598-020-73047-7. Sci Rep. 2020. PMID: 32985566 Free PMC article.
-
Epistasis Detection and Modeling for Genomic Selection in Cowpea (Vigna unguiculata L. Walp.).Front Genet. 2019 Jul 30;10:677. doi: 10.3389/fgene.2019.00677. eCollection 2019. Front Genet. 2019. PMID: 31417604 Free PMC article.
-
Effect of Low Temperature on Chlorophyll Biosynthesis and Chloroplast Biogenesis of Rice Seedlings during Greening.Int J Mol Sci. 2020 Feb 19;21(4):1390. doi: 10.3390/ijms21041390. Int J Mol Sci. 2020. PMID: 32092859 Free PMC article.
-
Substitution mapping of QTLs controlling seed dormancy using single segment substitution lines derived from multiple cultivated rice donors in seven cropping seasons.Theor Appl Genet. 2017 Jun;130(6):1191-1205. doi: 10.1007/s00122-017-2881-9. Epub 2017 Mar 10. Theor Appl Genet. 2017. PMID: 28283703
References
-
- Kobayashi Y. & Weigel D. Move on up, it's time for change--mobile signals controlling photoperiod-dependent flowering. Genes Dev 21, 2371–84 (2007). - PubMed
-
- Putterill J., Laurie R. & Macknight R. It's time to flower: the genetic control of flowering time. Bioessays 26, 363–73 (2004). - PubMed
-
- Tsuji H., Taoka K. & Shimamoto K. Regulation of flowering in rice: two florigen genes, a complex gene network, and natural variation. Curr Opin Plant Biol 14, 45–52 (2011). - PubMed
-
- Hayama R., Yokoi S., Tamaki S., Yano M. & Shimamoto K. Adaptation of photoperiodic control pathways produces short-day flowering in rice. Nature 422, 719–22 (2003). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

