Early specification of CD8+ T lymphocyte fates during adaptive immunity revealed by single-cell gene-expression analyses
- PMID: 24584088
- PMCID: PMC3968536
- DOI: 10.1038/ni.2842
Early specification of CD8+ T lymphocyte fates during adaptive immunity revealed by single-cell gene-expression analyses
Abstract
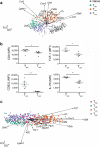
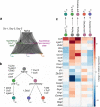
T lymphocytes responding to microbial infection give rise to effector cells that mediate acute host defense and memory cells that provide long-lived immunity, but the fundamental question of when and how these cells arise remains unresolved. Here we combined single-cell gene-expression analyses with 'machine-learning' approaches to trace the transcriptional 'roadmap' of individual CD8(+) T lymphocytes throughout the course of an immune response in vivo. Gene-expression signatures predictive of eventual fates could be discerned as early as the first T lymphocyte division and may have been influenced by asymmetric partitioning of the receptor for interleukin 2 (IL-2Rα) during mitosis. Our findings emphasize the importance of single-cell analyses in understanding fate determination and provide new insights into the specification of divergent lymphocyte fates early during an immune response to microbial infection.
Figures




Comment in
-
CD8(+) T cell diversification by asymmetric cell division.Nat Immunol. 2015 Sep;16(9):891-3. doi: 10.1038/ni.3235. Nat Immunol. 2015. PMID: 26287584 No abstract available.
-
Reply to: "CD8(+) T cell diversification by asymmetric cell division".Nat Immunol. 2015 Sep;16(9):893-4. doi: 10.1038/ni.3234. Nat Immunol. 2015. PMID: 26287585 Free PMC article. No abstract available.
References
-
- Ahmed R, Gray D. Immunological memory and protective immunity: understanding their relation. Science. 1996;272:54–60. - PubMed
-
- Masopust D, Kaech SM, Wherry EJ, Ahmed R. The role of programming in memory T-cell development. Curr Opin Immunol. 2004;16:217–25. - PubMed
-
- Sallusto F, Lenig D, Forster R, Lipp M, Lanzavecchia A. Two subsets of memory T lymphocytes with distinct homing potentials and effector functions. Nature. 1999;401:708–12. - PubMed
-
- Stemberger C, et al. A single naive CD8+ T cell precursor can develop into diverse effector and memory subsets. Immunity. 2007;27:985–97. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
- R24 DK080506/DK/NIDDK NIH HHS/United States
- DK80506/DK/NIDDK NIH HHS/United States
- K08 DK080949/DK/NIDDK NIH HHS/United States
- DK080949/DK/NIDDK NIH HHS/United States
- R01 HG004659/HG/NHGRI NIH HHS/United States
- P30 CA023100/CA/NCI NIH HHS/United States
- DP2 OD008469/OD/NIH HHS/United States
- R01 AI095277/AI/NIAID NIH HHS/United States
- HG004659/HG/NHGRI NIH HHS/United States
- HHMI/Howard Hughes Medical Institute/United States
- NS075449/NS/NINDS NIH HHS/United States
- OD008469/OD/NIH HHS/United States
- T32 DK007202/DK/NIDDK NIH HHS/United States
- AI095277/AI/NIAID NIH HHS/United States
- R01 NS075449/NS/NINDS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

