Supercoiling in DNA and chromatin
- PMID: 24584092
- PMCID: PMC4042020
- DOI: 10.1016/j.gde.2013.10.013
Supercoiling in DNA and chromatin
Abstract
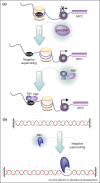
Supercoiling is a fundamental property of DNA and chromatin. It is modulated by polymerase and topoisomerase activities and, through regulated constraint, by DNA/chromatin binding proteins. As a non-covalent and elusive topological modification, supercoiling has proved intractable to research despite being a crucial regulator of nuclear structure and function. Recent studies have improved our understanding of the formation, regulation and organisation of supercoiling domains in vivo, and reinforce the prospect that the propagation of supercoiling can influence local and global chromatin structure. However, to further our understanding the development of new experimental tools and models are required to better dissect the mechanics of this key topological regulator.
Copyright © 2013 The Authors. Published by Elsevier Ltd.. All rights reserved.
Figures



Similar articles
-
Transcription forms and remodels supercoiling domains unfolding large-scale chromatin structures.Nat Struct Mol Biol. 2013 Mar;20(3):387-95. doi: 10.1038/nsmb.2509. Epub 2013 Feb 17. Nat Struct Mol Biol. 2013. PMID: 23416946 Free PMC article.
-
DNA Supercoiling, Topoisomerases, and Cohesin: Partners in Regulating Chromatin Architecture?Int J Mol Sci. 2018 Mar 16;19(3):884. doi: 10.3390/ijms19030884. Int J Mol Sci. 2018. PMID: 29547555 Free PMC article. Review.
-
Investigating DNA supercoiling in eukaryotic genomes.Brief Funct Genomics. 2017 Nov 1;16(6):379-389. doi: 10.1093/bfgp/elx007. Brief Funct Genomics. 2017. PMID: 28444308 Free PMC article. Review.
-
Synergistic Coordination of Chromatin Torsional Mechanics and Topoisomerase Activity.Cell. 2019 Oct 17;179(3):619-631.e15. doi: 10.1016/j.cell.2019.09.034. Cell. 2019. PMID: 31626768 Free PMC article.
-
Topoisomerase-modulated genome-wide DNA supercoiling domains colocalize with nuclear compartments and regulate human gene expression.Nat Struct Mol Biol. 2025 Jan;32(1):48-61. doi: 10.1038/s41594-024-01377-5. Epub 2024 Aug 16. Nat Struct Mol Biol. 2025. PMID: 39152238
Cited by
-
Simulated binding of transcription factors to active and inactive regions folds human chromosomes into loops, rosettes and topological domains.Nucleic Acids Res. 2016 May 5;44(8):3503-12. doi: 10.1093/nar/gkw135. Epub 2016 Apr 8. Nucleic Acids Res. 2016. PMID: 27060145 Free PMC article.
-
Small DNA circles as bacterial topoisomerase I inhibitors.RSC Adv. 2019 Jun 11;9(32):18415-18419. doi: 10.1039/c9ra02398d. eCollection 2019 Jun 10. RSC Adv. 2019. PMID: 35515216 Free PMC article.
-
Single-molecule micromanipulation studies of methylated DNA.Biophys J. 2021 Jun 1;120(11):2148-2155. doi: 10.1016/j.bpj.2021.03.039. Epub 2021 Apr 8. Biophys J. 2021. PMID: 33838135 Free PMC article.
-
The Tail That Wags the Dog: How the Disordered C-Terminal Domain Controls the Transcriptional Activities of the p53 Tumor-Suppressor Protein.Trends Biochem Sci. 2016 Dec;41(12):1022-1034. doi: 10.1016/j.tibs.2016.08.011. Epub 2016 Sep 23. Trends Biochem Sci. 2016. PMID: 27669647 Free PMC article. Review.
-
Impact of Chromosomal Architecture on the Function and Evolution of Bacterial Genomes.Front Microbiol. 2018 Aug 27;9:2019. doi: 10.3389/fmicb.2018.02019. eCollection 2018. Front Microbiol. 2018. PMID: 30210483 Free PMC article. Review.
References
-
- Freeman L.A.L., Garrard W.T.W. DNA supercoiling in chromatin structure and gene expression. Crit Rev Eukaryot Gene Expr. 1992;2:165–209. - PubMed
-
- Sinden R.R. Academic Press; 1994. DNA Structure and Function.
-
- Travers A.A., Muskhelishvili G.G. Bacterial chromatin. Curr Opin Genet Develop. 2005;15 8–8. - PubMed
-
- Norton V.G., Imai B.S., Yau P., Bradbury E.M. Histone acetylation reduces nucleosome core particle linking number change. Cell. 1989;57:449–457. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

