Fuzziness and noise in nucleosomal architecture
- PMID: 24586063
- PMCID: PMC4005669
- DOI: 10.1093/nar/gku165
Fuzziness and noise in nucleosomal architecture
Abstract
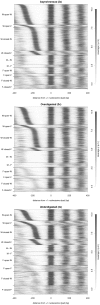
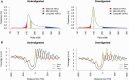
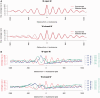
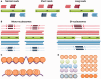
Nucleosome organization plays a key role in the regulation of gene expression. However, despite the striking advances in the accuracy of nucleosome maps, there are still severe discrepancies on individual nucleosome positioning and how this influences gene regulation. The variability among nucleosome maps, which precludes the fine analysis of nucleosome positioning, might emerge from diverse sources. We have carefully inspected the extrinsic factors that may induce diversity by the comparison of microccocal nuclease (MNase)-Seq derived nucleosome maps generated under distinct conditions. Furthermore, we have also explored the variation originated from intrinsic nucleosome dynamics by generating additional maps derived from cell cycle synchronized and asynchronous yeast cultures. Taken together, our study has enabled us to measure the effect of noise in nucleosome occupancy and positioning and provides insights into the underlying determinants. Furthermore, we present a systematic approach that may guide the standardization of MNase-Seq experiments in order to generate reproducible genome-wide nucleosome patterns.
Figures

Similar articles
-
Subtracting the sequence bias from partially digested MNase-seq data reveals a general contribution of TFIIS to nucleosome positioning.Epigenetics Chromatin. 2017 Dec 7;10(1):58. doi: 10.1186/s13072-017-0165-x. Epigenetics Chromatin. 2017. PMID: 29212533 Free PMC article.
-
The effect of micrococcal nuclease digestion on nucleosome positioning data.PLoS One. 2010 Dec 29;5(12):e15754. doi: 10.1371/journal.pone.0015754. PLoS One. 2010. PMID: 21206756 Free PMC article.
-
Application of MNase-Seq in the Global Mapping of Nucleosome Positioning in Plants.Methods Mol Biol. 2018;1830:353-366. doi: 10.1007/978-1-4939-8657-6_21. Methods Mol Biol. 2018. PMID: 30043381
-
Nucleosome positioning in yeasts: methods, maps, and mechanisms.Chromosoma. 2015 Jun;124(2):131-51. doi: 10.1007/s00412-014-0501-x. Epub 2014 Dec 23. Chromosoma. 2015. PMID: 25529773 Review.
-
Resolving nucleosomal positioning and occupancy with MNase-seq.Yi Chuan. 2020 Dec 17;42(12):1143-1155. doi: 10.16288/j.yczz.20-178. Yi Chuan. 2020. PMID: 33509779 Review.
Cited by
-
Autonomous actions of the human growth hormone long-range enhancer.Nucleic Acids Res. 2015 Feb 27;43(4):2091-101. doi: 10.1093/nar/gkv093. Epub 2015 Feb 6. Nucleic Acids Res. 2015. PMID: 25662214 Free PMC article.
-
The implication of DNA bending energy for nucleosome positioning and sliding.Sci Rep. 2018 Jun 11;8(1):8853. doi: 10.1038/s41598-018-27247-x. Sci Rep. 2018. PMID: 29891930 Free PMC article.
-
The PCNA-Pol δ complex couples lagging strand DNA synthesis to parental histone transfer for epigenetic inheritance.Sci Adv. 2024 Jun 7;10(23):eadn5175. doi: 10.1126/sciadv.adn5175. Epub 2024 Jun 5. Sci Adv. 2024. PMID: 38838138 Free PMC article.
-
Computational Approaches for Mining GRO-Seq Data to Identify and Characterize Active Enhancers.Methods Mol Biol. 2017;1468:121-38. doi: 10.1007/978-1-4939-4035-6_10. Methods Mol Biol. 2017. PMID: 27662874 Free PMC article.
-
An integrated machine-learning model to predict nucleosome architecture.Nucleic Acids Res. 2024 Sep 23;52(17):10132-10143. doi: 10.1093/nar/gkae689. Nucleic Acids Res. 2024. PMID: 39162225 Free PMC article.
References
-
- Kornberg RD, Lorch Y. Twenty-five years of the nucleosome, review fundamental particle of the eukaryote chromosome. Cell. 1999;98:285–294. - PubMed
-
- Malik HS, Henikoff S. Phylogenomics of the nucleosome. Nat. Struct. Biol. 2003;10:882–891. - PubMed
-
- Luger K, Mäder AW, Richmond RK, Sargent DF, Richmond TJ. Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature. 1997;389:251–260. - PubMed
-
- Tilgner H, Nikolaou C, Althammer S, Sammeth M, Beato M, Valcárcel J, Guigó R. Nucleosome positioning as a determinant of exon recognition. Nat. Struct. Mol. Biol. 2009;16:996–1001. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

