Selective disruption of aurora C kinase reveals distinct functions from aurora B kinase during meiosis in mouse oocytes
- PMID: 24586209
- PMCID: PMC3937256
- DOI: 10.1371/journal.pgen.1004194
Selective disruption of aurora C kinase reveals distinct functions from aurora B kinase during meiosis in mouse oocytes
Abstract
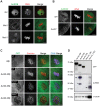
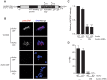
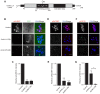
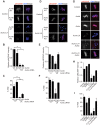
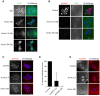
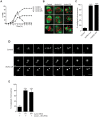
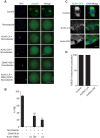
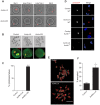
Aurora B kinase (AURKB) is the catalytic subunit of the chromosomal passenger complex (CPC), an essential regulator of chromosome segregation. In mitosis, the CPC is required to regulate kinetochore microtubule (K-MT) attachments, the spindle assembly checkpoint, and cytokinesis. Germ cells express an AURKB homolog, AURKC, which can also function in the CPC. Separation of AURKB and AURKC function during meiosis in oocytes by conventional approaches has not been successful. Therefore, the meiotic function of AURKC is still not fully understood. Here, we describe an ATP-binding-pocket-AURKC mutant, that when expressed in mouse oocytes specifically perturbs AURKC-CPC and not AURKB-CPC function. Using this mutant we show for the first time that AURKC has functions that do not overlap with AURKB. These functions include regulating localized CPC activity and regulating chromosome alignment and K-MT attachments at metaphase of meiosis I (Met I). We find that AURKC-CPC is not the sole CPC complex that regulates the spindle assembly checkpoint in meiosis, and as a result most AURKC-perturbed oocytes arrest at Met I. A small subset of oocytes do proceed through cytokinesis normally, suggesting that AURKC-CPC is not the sole CPC complex during telophase I. But, the resulting eggs are aneuploid, indicating that AURKC is a critical regulator of meiotic chromosome segregation in female gametes. Taken together, these data suggest that mammalian oocytes contain AURKC to efficiently execute meiosis I and ensure high-quality eggs necessary for sexual reproduction.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
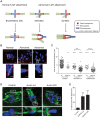
References
-
- Hassold T, Chiu D (1985) Maternal age-specific rates of numerical chromosome abnormalities with special reference to trisomy. Hum Genet 70: 11–17. - PubMed
-
- Hassold T, Hall H, Hunt P (2007) The origin of human aneuploidy: where we have been, where we are going. Hum Mol Genet 16 Spec No. 2: R203–208. - PubMed
-
- Hassold T, Hunt P (2001) To err (meiotically) is human: the genesis of human aneuploidy. Nat Rev Genet 2: 280–291. - PubMed
-
- Jones KT (2008) Meiosis in oocytes: predisposition to aneuploidy and its increased incidence with age. Hum Reprod Update 14: 143–158. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

