Mice carrying a hypomorphic Evi1 allele are embryonic viable but exhibit severe congenital heart defects
- PMID: 24586749
- PMCID: PMC3937339
- DOI: 10.1371/journal.pone.0089397
Mice carrying a hypomorphic Evi1 allele are embryonic viable but exhibit severe congenital heart defects
Abstract
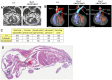
The ecotropic viral integration site 1 (Evi1) oncogenic transcription factor is one of a number of alternative transcripts encoded by the Mds1 and Evi1 complex locus (Mecom). Overexpression of Evi1 has been observed in a number of myeloid disorders and is associated with poor patient survival. It is also amplified and/or overexpressed in many epithelial cancers including nasopharyngeal carcinoma, ovarian carcinoma, ependymomas, and lung and colorectal cancers. Two murine knockout models have also demonstrated Evi1's critical role in the maintenance of hematopoietic stem cell renewal with its absence resulting in the death of mutant embryos due to hematopoietic failure. Here we characterize a novel mouse model (designated Evi1(fl3)) in which Evi1 exon 3, which carries the ATG start, is flanked by loxP sites. Unexpectedly, we found that germline deletion of exon3 produces a hypomorphic allele due to the use of an alternative ATG start site located in exon 4, resulting in a minor Evi1 N-terminal truncation and a block in expression of the Mds1-Evi1 fusion transcript. Evi1(δex3/δex3) mutant embryos showed only a mild non-lethal hematopoietic phenotype and bone marrow failure was only observed in adult Vav-iCre/+, Evi1(fl3/fl3) mice in which exon 3 was specifically deleted in the hematopoietic system. Evi1(δex3/δex3) knockout pups are born in normal numbers but die during the perinatal period from congenital heart defects. Database searches identified 143 genes with similar mutant heart phenotypes as those observed in Evi1(δex3/δex3) mutant pups. Interestingly, 42 of these congenital heart defect genes contain known Evi1-binding sites, and expression of 18 of these genes are also effected by Evi1 siRNA knockdown. These results show a potential functional involvement of Evi1 target genes in heart development and indicate that Evi1 is part of a transcriptional program that regulates cardiac development in addition to the development of blood.
Conflict of interest statement
Figures






Similar articles
-
Functional features of EVI1 and EVI1Δ324 isoforms of MECOM gene in genome-wide transcription regulation and oncogenicity.Oncogene. 2016 May 5;35(18):2311-21. doi: 10.1038/onc.2015.286. Epub 2015 Aug 3. Oncogene. 2016. PMID: 26234679
-
Oncogenic transcription factor Evi1 regulates hematopoietic stem cell proliferation through GATA-2 expression.EMBO J. 2005 Jun 1;24(11):1976-87. doi: 10.1038/sj.emboj.7600679. Epub 2005 May 12. EMBO J. 2005. PMID: 15889140 Free PMC article.
-
[The role and regulation of EVI1 in normal hematopoiesis and hematopoietic malignancies].Rinsho Ketsueki. 2024;65(9):954-960. doi: 10.11406/rinketsu.65.954. Rinsho Ketsueki. 2024. PMID: 39358295 Review. Japanese.
-
Mutations in MECOM, Encoding Oncoprotein EVI1, Cause Radioulnar Synostosis with Amegakaryocytic Thrombocytopenia.Am J Hum Genet. 2015 Dec 3;97(6):848-54. doi: 10.1016/j.ajhg.2015.10.010. Epub 2015 Nov 12. Am J Hum Genet. 2015. PMID: 26581901 Free PMC article.
-
The role of EVI1 in normal and leukemic cells.Blood Cells Mol Dis. 2003 Sep-Oct;31(2):206-12. doi: 10.1016/s1079-9796(03)00159-1. Blood Cells Mol Dis. 2003. PMID: 12972028 Review.
Cited by
-
Molecular Pathways and Animal Models of Truncus Arteriosus.Adv Exp Med Biol. 2024;1441:853-865. doi: 10.1007/978-3-031-44087-8_52. Adv Exp Med Biol. 2024. PMID: 38884754 Review.
-
SOX9 in organogenesis: shared and unique transcriptional functions.Cell Mol Life Sci. 2022 Sep 17;79(10):522. doi: 10.1007/s00018-022-04543-4. Cell Mol Life Sci. 2022. PMID: 36114905 Free PMC article. Review.
-
The Roles of Histone Lysine Methyltransferases in Heart Development and Disease.J Cardiovasc Dev Dis. 2023 Jul 18;10(7):305. doi: 10.3390/jcdd10070305. J Cardiovasc Dev Dis. 2023. PMID: 37504561 Free PMC article. Review.
-
Mecom mutation related to radioulnar synostosis with amegakaryocytic thrombocytopenia reduces HSPCs in mice.Blood Adv. 2023 Sep 26;7(18):5409-5420. doi: 10.1182/bloodadvances.2022008462. Blood Adv. 2023. PMID: 37099686 Free PMC article.
-
Functional features of EVI1 and EVI1Δ324 isoforms of MECOM gene in genome-wide transcription regulation and oncogenicity.Oncogene. 2016 May 5;35(18):2311-21. doi: 10.1038/onc.2015.286. Epub 2015 Aug 3. Oncogene. 2016. PMID: 26234679
References
-
- Nerlov C (2010) Transcriptional and translational control of C/EBPs: the case for “deep” genetics to understand physiological function. Bioessays 32: 680–686. - PubMed
-
- Lugthart S, van Drunen E, van Norden Y, van Hoven A, Erpelinck CA, et al. (2008) High EVI1 levels predict adverse outcome in acute myeloid leukemia: prevalence of EVI1 overexpression and chromosome 3q26 abnormalities underestimated. Blood 111: 4329–4337. - PubMed
-
- Ogawa S, Kurokawa M, Tanaka T, Mitani K, Inazawa J, et al. (1996) Structurally altered Evi-1 protein generated in the 3q21q26 syndrome. Oncogene 13: 183–191. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

