Insight into highly conserved H1 subtype-specific epitopes in influenza virus hemagglutinin
- PMID: 24587046
- PMCID: PMC3935945
- DOI: 10.1371/journal.pone.0089803
Insight into highly conserved H1 subtype-specific epitopes in influenza virus hemagglutinin
Abstract
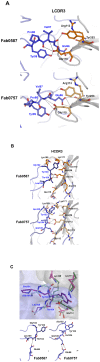
Influenza viruses continuously undergo antigenic changes with gradual accumulation of mutations in hemagglutinin (HA) that is a major determinant in subtype specificity. The identification of conserved epitopes within specific HA subtypes gives an important clue for developing new vaccines and diagnostics. We produced and characterized nine monoclonal antibodies that showed significant neutralizing activities against H1 subtype influenza viruses, and determined the complex structure of HA derived from a 2009 pandemic virus A/Korea/01/2009 (KR01) and the Fab fragment from H1-specific monoclonal antibody GC0587. The overall structure of the complex was essentially identical to the previously determined KR01 HA-Fab0757 complex structure. Both Fab0587 and Fab0757 recognize readily accessible head regions of HA, revealing broadly shared and conserved antigenic determinants among H1 subtypes. The β-strands constituted by Ser110-Glu115 and Lys169-Lys170 form H1 epitopes with distinct conformations from those of H1 and H3 HA sites. In particular, Glu112, Glu115, Lys169, and Lys171 that are highly conserved among H1 subtype HAs have close contacts with HCDR3 and LCDR3. The differences between Fab0587 and Fab0757 complexes reside mainly in HCDR3 and LCDR3, providing distinct antigenic determinants specific for 1918 pdm influenza strain. Our results demonstrate a potential key neutralizing epitope important for H1 subtype specificity in influenza virus.
Conflict of interest statement
Figures
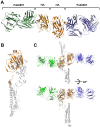
Similar articles
-
Alternative recognition of the conserved stem epitope in influenza A virus hemagglutinin by a VH3-30-encoded heterosubtypic antibody.J Virol. 2014 Jun;88(12):7083-92. doi: 10.1128/JVI.00178-14. Epub 2014 Apr 9. J Virol. 2014. PMID: 24719426 Free PMC article.
-
Mapping of a Novel H3-Specific Broadly Neutralizing Monoclonal Antibody Targeting the Hemagglutinin Globular Head Isolated from an Elite Influenza Virus-Immunized Donor Exhibiting Serological Breadth.J Virol. 2020 Feb 28;94(6):e01035-19. doi: 10.1128/JVI.01035-19. Print 2020 Feb 28. J Virol. 2020. PMID: 31826999 Free PMC article.
-
A broadly neutralizing human monoclonal antibody directed against a novel conserved epitope on the influenza virus H3 hemagglutinin globular head.J Virol. 2014 Jun;88(12):6743-50. doi: 10.1128/JVI.03562-13. Epub 2014 Apr 2. J Virol. 2014. PMID: 24696468 Free PMC article.
-
The antigenic architecture of the hemagglutinin of influenza H5N1 viruses.Mol Immunol. 2013 Dec;56(4):705-19. doi: 10.1016/j.molimm.2013.07.010. Epub 2013 Aug 7. Mol Immunol. 2013. PMID: 23933511 Review.
-
Neutralizing Antibodies Targeting the Conserved Stem Region of Influenza Hemagglutinin.Vaccines (Basel). 2020 Jul 12;8(3):382. doi: 10.3390/vaccines8030382. Vaccines (Basel). 2020. PMID: 32664628 Free PMC article. Review.
Cited by
-
A Highly Potent and Broadly Neutralizing H1 Influenza-Specific Human Monoclonal Antibody.Sci Rep. 2018 Mar 12;8(1):4374. doi: 10.1038/s41598-018-22307-8. Sci Rep. 2018. PMID: 29531320 Free PMC article.
-
Influenza virus antigenicity and broadly neutralizing epitopes.Curr Opin Virol. 2015 Apr;11:113-21. doi: 10.1016/j.coviro.2015.03.006. Epub 2015 Apr 4. Curr Opin Virol. 2015. PMID: 25846699 Free PMC article. Review.
-
Comparison of the Protective Efficacy of Neutralizing Epitopes of 2009 Pandemic H1N1 Influenza Hemagglutinin.Front Immunol. 2017 Aug 31;8:1070. doi: 10.3389/fimmu.2017.01070. eCollection 2017. Front Immunol. 2017. PMID: 28912784 Free PMC article.
-
Rapid affinity optimization of an anti-TREM2 clinical lead antibody by cross-lineage immune repertoire mining.Nat Commun. 2024 Sep 27;15(1):8382. doi: 10.1038/s41467-024-52442-y. Nat Commun. 2024. PMID: 39333507 Free PMC article.
-
Restricted epitope specificity determined by variable region germline segment pairing in rodent antibody repertoires.MAbs. 2020 Jan-Dec;12(1):1722541. doi: 10.1080/19420862.2020.1722541. MAbs. 2020. PMID: 32041466 Free PMC article.
References
-
- Dushoff J, Plotkin JB, Viboud C, Earn DJ, Simonsen L (2006) Mortality due to influenza in the United States–an annualized regression approach using multiple-cause mortality data. Am J Epidemiol 163: 181–187. - PubMed
-
- Johnson NP, Mueller J (2002) Updating the accounts: global mortality of the 1918–1920 “Spanish” influenza pandemic. Bull Hist Med 76: 105–115. - PubMed
-
- WHO.
-
- Cox NJ, Subbarao K (2000) Global epidemiology of influenza: past and present. Annu Rev Med 51: 407–421. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources

