In search of novel drug target sites on estrogen receptors using RNA aptamers
- PMID: 24588102
- PMCID: PMC4026216
- DOI: 10.1089/nat.2013.0474
In search of novel drug target sites on estrogen receptors using RNA aptamers
Abstract
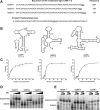
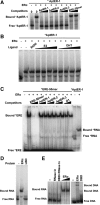
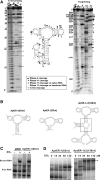
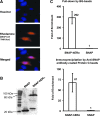
Estrogen receptor α (ERα) is a well-validated drug target for a majority of breast cancers. But the target sites on this receptor are far from exhaustively defined. Almost all ER antagonists in clinical use function by binding to the ligand-binding pocket to occlude agonist access. Resistance to this type of drugs may develop over time, not caused by the change of ERα itself, but by changes in ER associated proteins. This observation is fueling the development of reagents that downregulate ER activity through novel binding sites. However, it is challenging to find general ER antagonists that act independently from other known ER ligands. In this report, we describe the utility of RNA aptamers in the search for new drug target sites on ERα. We have identified three high affinity aptamers and characterized one of them in detail. This aptamer interacted with ERα in a way not affected by the presence or absence of either the steroidal ligands or the estrogen response DNA elements, and effectively inhibited ER-mediated transcriptional activation in a breast cancer cell line. Serving as a novel drug lead, it may also be used to guide the rational chemical synthesis of small molecule drugs or to perform screens of small molecule libraries for those that are able to displace the aptamer from its binding site.
Figures
Similar articles
-
Aptamer-Assisted Detection of the Altered Expression of Estrogen Receptor Alpha in Human Breast Cancer.PLoS One. 2016 Apr 4;11(4):e0153001. doi: 10.1371/journal.pone.0153001. eCollection 2016. PLoS One. 2016. PMID: 27043307 Free PMC article.
-
In silico selection of an aptamer to estrogen receptor alpha using computational docking employing estrogen response elements as aptamer-alike molecules.Sci Rep. 2016 Feb 22;6:21285. doi: 10.1038/srep21285. Sci Rep. 2016. PMID: 26899418 Free PMC article.
-
In silico identification of novel inhibitors targeting the DNA-binding domain of the human estrogen receptor alpha.J Steroid Biochem Mol Biol. 2021 Oct;213:105966. doi: 10.1016/j.jsbmb.2021.105966. Epub 2021 Aug 17. J Steroid Biochem Mol Biol. 2021. PMID: 34416373
-
Molecular mechanisms of estrogen action: selective ligands and receptor pharmacology.J Steroid Biochem Mol Biol. 2000 Nov 30;74(5):279-85. doi: 10.1016/s0960-0760(00)00104-7. J Steroid Biochem Mol Biol. 2000. PMID: 11162936 Review.
-
Computer-Aided Ligand Discovery for Estrogen Receptor Alpha.Int J Mol Sci. 2020 Jun 12;21(12):4193. doi: 10.3390/ijms21124193. Int J Mol Sci. 2020. PMID: 32545494 Free PMC article. Review.
Cited by
-
Aptamer-Enabled Manipulation of the Hsp70 Chaperone System Suggests a Novel Strategy for Targeted Ubiquitination.Nucleic Acid Ther. 2016 Feb;26(1):20-8. doi: 10.1089/nat.2015.0563. Epub 2015 Dec 7. Nucleic Acid Ther. 2016. PMID: 26640962 Free PMC article.
-
Selection of DNA aptamers for extra cellular domain of human epidermal growth factor receptor 2 to detect HER2 positive carcinomas.Clin Transl Oncol. 2017 Aug;19(8):976-988. doi: 10.1007/s12094-017-1629-y. Epub 2017 Feb 21. Clin Transl Oncol. 2017. PMID: 28224267
-
Aptamer-Assisted Detection of the Altered Expression of Estrogen Receptor Alpha in Human Breast Cancer.PLoS One. 2016 Apr 4;11(4):e0153001. doi: 10.1371/journal.pone.0153001. eCollection 2016. PLoS One. 2016. PMID: 27043307 Free PMC article.
-
An RNA aptamer specific to Hsp70-ATP conformation inhibits its ATPase activity independent of Hsp40.Nucleic Acid Ther. 2015 Apr;25(2):103-12. doi: 10.1089/nat.2014.0510. Epub 2015 Feb 5. Nucleic Acid Ther. 2015. PMID: 25654640 Free PMC article.
-
In silico selection of an aptamer to estrogen receptor alpha using computational docking employing estrogen response elements as aptamer-alike molecules.Sci Rep. 2016 Feb 22;6:21285. doi: 10.1038/srep21285. Sci Rep. 2016. PMID: 26899418 Free PMC article.
References
-
- BAN N., NISSEN P., HANSEN J., MOORE P.B., and STEITZ T.A. (2000). The complete atomic structure of the large ribosomal subunit at 2.4 A resolution. Science 289,905–920 - PubMed
-
- CARRAZ M., ZWART W., PHAN T., MICHALIDES R., and BRUNSVELD L. (2009). Perturbation of estrogen receptor alpha localization with synthetic nona-arginine LXXLL-peptide coactivator binding inhibitors. Chem. Biol. 16,702–711 - PubMed
-
- CHANG C., NORRIS J.D., GRON H., PAIGE L.A., HAMILTON P.T., KENAN D.J., FOWLKES D., and MCDONNELL D.P. (1999). Dissection of the LXXLL nuclear receptor-coactivator interaction motif using combinatorial peptide libraries: discovery of peptide antagonists of estrogen receptors alpha and beta. Mol. Cell Biol. 19,8226–8239 - PMC - PubMed
-
- CIRILLO F., NASSA G., TARALLO R., STELLATO C., DE FILIPPO M.R., AMBROSINO C., BAUMANN M., NYMAN T.A., and WEISZ A. (2013). Molecular mechanisms of selective estrogen receptor modulator activity in human breast cancer cells: identification of novel nuclear cofactors of antiestrogen-ERalpha complexes by interaction proteomics. J. Proteome Res. 12,421–431 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

