Meiosis-specific cohesin mediates homolog recognition in mouse spermatocytes
- PMID: 24589552
- PMCID: PMC3967048
- DOI: 10.1101/gad.237313.113
Meiosis-specific cohesin mediates homolog recognition in mouse spermatocytes
Abstract
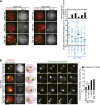
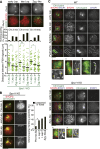
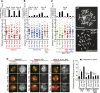
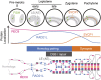
During meiosis, homologous chromosome (homolog) pairing is promoted by several layers of regulation that include dynamic chromosome movement and meiotic recombination. However, the way in which homologs recognize each other remains a fundamental issue in chromosome biology. Here, we show that homolog recognition or association initiates upon entry into meiotic prophase before axis assembly and double-strand break (DSB) formation. This homolog association develops into tight pairing only during or after axis formation. Intriguingly, the ability to recognize homologs is retained in Sun1 knockout spermatocytes, in which telomere-directed chromosome movement is abolished, and this is the case even in Spo11 knockout spermatocytes, in which DSB-dependent DNA homology search is absent. Disruption of meiosis-specific cohesin RAD21L precludes the initial association of homologs as well as the subsequent pairing in spermatocytes. These findings suggest the intriguing possibility that homolog recognition is achieved primarily by searching for homology in the chromosome architecture as defined by meiosis-specific cohesin rather than in the DNA sequence itself.
Keywords: DSB; bouquet; cohesin; homolog pairing.
Figures



Comment in
-
Which one is the real matchmaker for the pair?Asian J Androl. 2014 Sep-Oct;16(5):667-8. doi: 10.4103/1008-682X.133316. Asian J Androl. 2014. PMID: 25038183 Free PMC article.
References
-
- Bannister LA, Reinholdt LG, Munroe RJ, Schimenti JC 2004. Positional cloning and characterization of mouse mei8, a disrupted allelle of the meiotic cohesin Rec8. Genesis 40: 184–194 - PubMed
-
- Barzel A, Kupiec M 2008. Finding a match: How do homologous sequences get together for recombination? Nat Rev Genet 9: 27–37 - PubMed
-
- Baudat F, de Massy B 2007. Regulating double-stranded DNA break repair towards crossover or non-crossover during mammalian meiosis. Chromosome Res 15: 565–577 - PubMed
-
- Baudat F, Manova K, Yuen JP, Jasin M, Keeney S 2000. Chromosome synapsis defects and sexually dimorphic meiotic progression in mice lacking Spo11. Mol Cell 6: 989–998 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous
