Innate DNA sensing is impaired in HIV patients and IFI16 expression correlates with chronic immune activation
- PMID: 24593816
- PMCID: PMC4089180
- DOI: 10.1111/cei.12317
Innate DNA sensing is impaired in HIV patients and IFI16 expression correlates with chronic immune activation
Abstract
The innate immune system has been recognized to play a role in the pathogenesis of HIV infection, both by stimulating protective activities and through a contribution to chronic immune activation, the development of immunodeficiency and progression to AIDS. A role for DNA sensors in HIV recognition has been suggested recently, and the aim of the present study was to describe the influence of HIV infection on expression and function of intracellular DNA sensing. Here we demonstrate impaired expression of interferon-stimulated genes in responses to DNA in peripheral blood monuclear cells from HIV-positive individuals, irrespective of whether patients receive anti-retroviral treatment. Furthermore, we show that expression levels of the DNA sensors interferon-inducible protein 16 (IFI16) and cyclic guanosine monophosphate-adenosine monophosphate synthase were increased in treatment-naive patients, and for IFI16 expression was correlated with high viral load and low CD4 cell count. Finally, our data demonstrate a correlation between IFI16 and CD38 expression, a marker of immune activation, in CD4(+) central and effector memory T cells, which may indicate that IFI16-mediated DNA sensing and signalling contributes to chronic immune activation. Altogether, the present study demonstrates abnormal expression and function of cytosolic DNA sensors in HIV patients, which may have implications for control of opportunistic infections, chronic immune activation and T cell death.
Keywords: DNA sensing; HIV; IFI16; cGAS; immune activation; interferon-stimulated genes.
© 2014 British Society for Immunology.
Figures



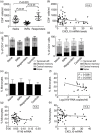

References
-
- Barre-Sinoussi F, Chermann JC, Rey F, et al. Isolation of a T-lymphotropic retrovirus from a patient at risk for acquired immune deficiency syndrome (AIDS) Science. 1983;220:868–871. - PubMed
-
- De Cock KM, Jaffe HW, Curran JW. The evolving epidemiology of HIV/AIDS. AIDS. 2012;26:1205–1213. - PubMed
-
- Mayer K, Beyrer C. WHO's new HIV guidelines: opportunities and challenges. Lancet. 2013;382:287–288. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

