Nucleation precursors in protein crystallization
- PMID: 24598910
- PMCID: PMC3944685
- DOI: 10.1107/S2053230X14002386
Nucleation precursors in protein crystallization
Abstract
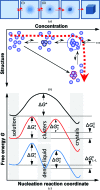
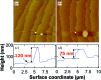
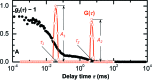
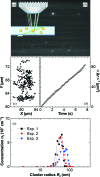
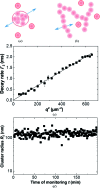
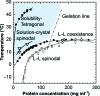
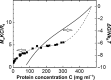
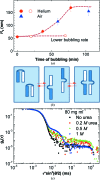
Protein crystal nucleation is a central problem in biological crystallography and other areas of science, technology and medicine. Recent studies have demonstrated that protein crystal nuclei form within crucial precursors. Here, methods of detection and characterization of the precursors are reviewed: dynamic light scattering, atomic force microscopy and Brownian microscopy. Data for several proteins provided by these methods have demonstrated that the nucleation precursors are clusters consisting of protein-dense liquid, which are metastable with respect to the host protein solution. The clusters are several hundred nanometres in size, the cluster population occupies from 10(-7) to 10(-3) of the solution volume, and their properties in solutions supersaturated with respect to crystals are similar to those in homogeneous, i.e. undersaturated, solutions. The clusters exist owing to the conformation flexibility of the protein molecules, leading to exposure of hydrophobic surfaces and enhanced intermolecular binding. These results indicate that protein conformational flexibility might be the mechanism behind the metastable mesoscopic clusters and crystal nucleation. Investigations of the cluster properties are still in their infancy. Results on direct imaging of cluster behaviors and characterization of cluster mechanisms with a variety of proteins will soon lead to major breakthroughs in protein biophysics.
Keywords: crystallization; nucleation mechanism; partial unfolding; protein crystals; protein-rich clusters; two-step nucleation.
Figures

Similar articles
-
Do protein crystals nucleate within dense liquid clusters?Acta Crystallogr F Struct Biol Commun. 2015 Jul;71(Pt 7):815-22. doi: 10.1107/S2053230X15008997. Epub 2015 Jun 27. Acta Crystallogr F Struct Biol Commun. 2015. PMID: 26144225 Free PMC article.
-
Metastable liquid clusters in super- and undersaturated protein solutions.J Phys Chem B. 2007 Mar 29;111(12):3106-14. doi: 10.1021/jp068827o. Epub 2007 Mar 8. J Phys Chem B. 2007. PMID: 17388477
-
Recent advances in the understanding of two-step nucleation of protein crystals.Faraday Discuss. 2015;179:27-40. doi: 10.1039/c4fd00217b. Epub 2015 Apr 10. Faraday Discuss. 2015. PMID: 25859918
-
Nonclassical nucleation pathways in protein crystallization.J Phys Condens Matter. 2017 Nov 8;29(44):443002. doi: 10.1088/1361-648X/aa8253. J Phys Condens Matter. 2017. PMID: 28984274 Review.
-
Phase diagrams and kinetics of phase transitions in protein solutions.J Phys Condens Matter. 2012 May 16;24(19):193101. doi: 10.1088/0953-8984/24/19/193101. Epub 2012 Apr 11. J Phys Condens Matter. 2012. PMID: 22495288 Review.
Cited by
-
Do protein crystals nucleate within dense liquid clusters?Acta Crystallogr F Struct Biol Commun. 2015 Jul;71(Pt 7):815-22. doi: 10.1107/S2053230X15008997. Epub 2015 Jun 27. Acta Crystallogr F Struct Biol Commun. 2015. PMID: 26144225 Free PMC article.
-
Tracking Prenucleation Molecular Clustering of Salicylamide in Organic Solvents.Cryst Growth Des. 2024 Jun 21;24(13):5740-5753. doi: 10.1021/acs.cgd.4c00507. eCollection 2024 Jul 3. Cryst Growth Des. 2024. PMID: 38983120 Free PMC article.
-
The "Sticky Patch" Model of Crystallization and Modification of Proteins for Enhanced Crystallizability.Methods Mol Biol. 2017;1607:77-115. doi: 10.1007/978-1-4939-7000-1_4. Methods Mol Biol. 2017. PMID: 28573570 Free PMC article. Review.
-
Automation in biological crystallization.Acta Crystallogr F Struct Biol Commun. 2014 Jun;70(Pt 6):686-96. doi: 10.1107/S2053230X14011601. Epub 2014 May 28. Acta Crystallogr F Struct Biol Commun. 2014. PMID: 24915074 Free PMC article.
-
Liquid Metastable Precursors of Ibuprofen as Aqueous Nucleation Intermediates.Angew Chem Int Ed Engl. 2019 Dec 19;58(52):19103-19109. doi: 10.1002/anie.201910986. Epub 2019 Nov 6. Angew Chem Int Ed Engl. 2019. PMID: 31556970 Free PMC article.
References
-
- Aber, J. E., Arnold, S. & Garetz, B. A. (2005). Strong DC Electric Field Applied to Supersaturated Aqueous Glycine Solution Induces Nucleation of the Polymorph. Phys. Rev. Lett. 94, 145503. - PubMed
-
- Asherie, N., Lomakin, A. & Benedek, G. B. (1996). Phase Diagram of Colloidal Solutions. Phys. Rev. Lett. 77, 4832–4835. - PubMed
-
- Atkins, P. & DePaula, J. (2002). Physical Chemistry, 7th ed. New York: W. H. Freeman & Co.
-
- Auer, S. & Frenkel, D. (2004). Numerical prediction of absolute crystallization rates in hard-sphere colloids. J. Chem. Phys. 120, 3015–3029. - PubMed
-
- Bekard, I. B., Asimakis, P., Bertolini, J. & Dunstan, D. E. (2011). The effects of shear flow on protein structure and function. Biopolymers, 95, 733–745. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

