Deubiquitylating enzymes and their emerging role in plant biology
- PMID: 24600466
- PMCID: PMC3928566
- DOI: 10.3389/fpls.2014.00056
Deubiquitylating enzymes and their emerging role in plant biology
Abstract
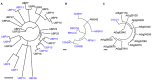
Ubiquitylation is a reversible post-translational modification that is involved in various cellular pathways and that thereby regulates various aspects of plant biology. For a long time, functional studies of ubiquitylation have focused on the function of ubiquitylating enzymes, especially the E3 ligases, rather than deubiquitylating enzymes (DUBs) or ubiquitin isopeptidases, enzymes that hydrolyze ubiquitin chains. One reason may be the smaller number of DUBs in comparison to E3 ligases, implying the broader substrate specificities of DUBs and the difficulties to identify the direct targets. However, recent studies have revealed that DUBs also actively participate in controlling cellular events and thus play pivotal roles in plant development and growth. DUBs are also essential for processing ubiquitin precursors and are important for recycling ubiquitin molecules from target proteins prior to their degradation and thereby maintaining the free ubiquitin pool in the cell. Here, we will discuss the five different DUB families (USP/UBP, UCH, JAMM, OTU, and MJD) and their known biochemical and physiological roles in plants.
Keywords: DUB; JAMM; MJD; OTU; UBP; UCH; USP; deubiquitylation.
Figures
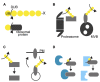
Similar articles
-
Monitoring Target Engagement of Deubiquitylating Enzymes Using Activity Probes: Past, Present, and Future.Methods Mol Biol. 2016;1449:395-410. doi: 10.1007/978-1-4939-3756-1_26. Methods Mol Biol. 2016. PMID: 27613052 Free PMC article.
-
Plant deubiquitinases: from structure and activity to biological functions.Plant Cell Rep. 2023 Mar;42(3):469-486. doi: 10.1007/s00299-022-02962-y. Epub 2022 Dec 26. Plant Cell Rep. 2023. PMID: 36567335 Review.
-
Deubiquitylating enzymes in Arabidopsis thaliana endocytic protein degradation.Biochem Soc Trans. 2024 Feb 28;52(1):291-299. doi: 10.1042/BST20230561. Biochem Soc Trans. 2024. PMID: 38174770 Review.
-
Deubiquitinases in skeletal muscle-the underappreciated side of the ubiquitination coin.Am J Physiol Cell Physiol. 2024 Dec 1;327(6):C1651-C1665. doi: 10.1152/ajpcell.00553.2024. Epub 2024 Sep 30. Am J Physiol Cell Physiol. 2024. PMID: 39344415 Review.
-
Erasing marks: Functions of plant deubiquitylating enzymes in modulating the ubiquitin code.Plant Cell. 2024 Sep 3;36(9):3057-3073. doi: 10.1093/plcell/koae129. Plant Cell. 2024. PMID: 38656977 Free PMC article. Review.
Cited by
-
Lipid-mediated activation of plasma membrane-localized deubiquitylating enzymes modulate endosomal trafficking.Nat Commun. 2022 Nov 12;13(1):6897. doi: 10.1038/s41467-022-34637-3. Nat Commun. 2022. PMID: 36371501 Free PMC article.
-
The Beginning of the End: Initial Steps in the Degradation of Plasma Membrane Proteins.Front Plant Sci. 2020 May 21;11:680. doi: 10.3389/fpls.2020.00680. eCollection 2020. Front Plant Sci. 2020. PMID: 32528512 Free PMC article. Review.
-
A dynamic ubiquitination balance of cell proliferation and endoreduplication regulators determines plant organ size.Sci Adv. 2024 Mar 15;10(11):eadj2570. doi: 10.1126/sciadv.adj2570. Epub 2024 Mar 13. Sci Adv. 2024. PMID: 38478622 Free PMC article.
-
Ubiquitination of the ubiquitin-binding machinery: how early ESCRT components are controlled.Essays Biochem. 2022 Aug 5;66(2):169-177. doi: 10.1042/EBC20210042. Essays Biochem. 2022. PMID: 35352804 Free PMC article. Review.
-
The deubiquitinases UBP12 and UBP13 integrate with the E3 ubiquitin ligase XBAT35.2 to modulate VPS23A stability in ABA signaling.Sci Adv. 2022 Apr 8;8(14):eabl5765. doi: 10.1126/sciadv.abl5765. Epub 2022 Apr 6. Sci Adv. 2022. PMID: 35385312 Free PMC article.
References
-
- Callis J., Raasch J. A., Vierstra R. D. (1990). Ubiquitin extension proteins of Arabidopsis thaliana. Structure, localization, and expression of their promoters in transgenic tobacco J. Biol. Chem. 265 12486–12493 - PubMed
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources

