Nucleosome acidic patch promotes RNF168- and RING1B/BMI1-dependent H2AX and H2A ubiquitination and DNA damage signaling
- PMID: 24603765
- PMCID: PMC3945288
- DOI: 10.1371/journal.pgen.1004178
Nucleosome acidic patch promotes RNF168- and RING1B/BMI1-dependent H2AX and H2A ubiquitination and DNA damage signaling
Abstract
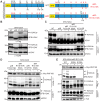
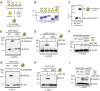
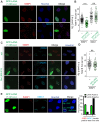
Histone ubiquitinations are critical for the activation of the DNA damage response (DDR). In particular, RNF168 and RING1B/BMI1 function in the DDR by ubiquitinating H2A/H2AX on Lys-13/15 and Lys-118/119, respectively. However, it remains to be defined how the ubiquitin pathway engages chromatin to provide regulation of ubiquitin targeting of specific histone residues. Here we identify the nucleosome acid patch as a critical chromatin mediator of H2A/H2AX ubiquitination (ub). The acidic patch is required for RNF168- and RING1B/BMI1-dependent H2A/H2AXub in vivo. The acidic patch functions within the nucleosome as nucleosomes containing a mutated acidic patch exhibit defective H2A/H2AXub by RNF168 and RING1B/BMI1 in vitro. Furthermore, direct perturbation of the nucleosome acidic patch in vivo by the expression of an engineered acidic patch interacting viral peptide, LANA, results in defective H2AXub and RNF168-dependent DNA damage responses including 53BP1 and BRCA1 recruitment to DNA damage. The acidic patch therefore is a critical nucleosome feature that may serve as a scaffold to integrate multiple ubiquitin signals on chromatin to compose selective ubiquitinations on histones for DNA damage signaling.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures




References
-
- Kouzarides T (2007) Chromatin modifications and their function. Cell 128: 693–705. - PubMed
-
- Miller KM, Jackson SP (2012) Histone marks: repairing DNA breaks within the context of chromatin. Biochem Soc Trans 40: 370–376. - PubMed
-
- Rogakou EP, Pilch DR, Orr AH, Ivanova VS, Bonner WM (1998) DNA double-stranded breaks induce histone H2AX phosphorylation on serine 139. J Biol Chem 273: 5858–5868. - PubMed
-
- Paull TT, Rogakou EP, Yamazaki V, Kirchgessner CU, Gellert M, et al. (2000) A critical role for histone H2AX in recruitment of repair factors to nuclear foci after DNA damage. Curr Biol 10: 886–895. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

