Structural basis for heavy metal detoxification by an Atm1-type ABC exporter
- PMID: 24604198
- PMCID: PMC4151877
- DOI: 10.1126/science.1246489
Structural basis for heavy metal detoxification by an Atm1-type ABC exporter
Abstract
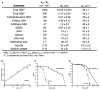
Although substantial progress has been achieved in the structural analysis of exporters from the superfamily of adenosine triphosphate (ATP)-binding cassette (ABC) transporters, much less is known about how they selectively recognize substrates and how substrate binding is coupled to ATP hydrolysis. We have addressed these questions through crystallographic analysis of the Atm1/ABCB7/HMT1/ABCB6 ortholog from Novosphingobium aromaticivorans DSM 12444, NaAtm1, at 2.4 angstrom resolution. Consistent with a physiological role in cellular detoxification processes, functional studies showed that glutathione derivatives can serve as substrates for NaAtm1 and that its overexpression in Escherichia coli confers protection against silver and mercury toxicity. The glutathione binding site highlights the articulated design of ABC exporters, with ligands and nucleotides spanning structurally conserved elements to create adaptable interfaces accommodating conformational rearrangements during the transport cycle.
Figures
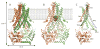


References
-
- Lill R, Kispal G. Res Microbiol. 2001;152:331. - PubMed
-
- Holland IB, Cole SPC, Kuchler K, Higgins CF. ABC proteins: From Bacteria to Man. Academic Press; London: 2003. p. 647.
-
- Zutz A, Gompf S, Schagger H, Tampé R. Biochim Biophys Acta. 2009;1787:681. - PubMed
-
- Lill R, et al. Biochim Biophys Acta. 2012;1823:1491. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

