Six-color intravital two-photon imaging of brain tumors and their dynamic microenvironment
- PMID: 24605087
- PMCID: PMC3932518
- DOI: 10.3389/fncel.2014.00057
Six-color intravital two-photon imaging of brain tumors and their dynamic microenvironment
Abstract
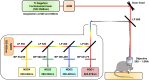
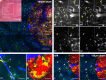
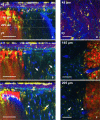
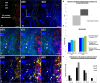
The majority of intravital studies on brain tumor in living animal so far rely on dual color imaging. We describe here a multiphoton imaging protocol to dynamically characterize the interactions between six cellular components in a living mouse. We applied this methodology to a clinically relevant glioblastoma multiforme (GBM) model designed in reporter mice with targeted cell populations labeled by fluorescent proteins of different colors. This model permitted us to make non-invasive longitudinal and multi-scale observations of cell-to-cell interactions. We provide examples of such 5D (x,y,z,t,color) images acquired on a daily basis from volumes of interest, covering most of the mouse parietal cortex at subcellular resolution. Spectral deconvolution allowed us to accurately separate each cell population as well as some components of the extracellular matrix. The technique represents a powerful tool for investigating how tumor progression is influenced by the interactions of tumor cells with host cells and the extracellular matrix micro-environment. It will be especially valuable for evaluating neuro-oncological drug efficacy and target specificity. The imaging protocol provided here can be easily translated to other mouse models of neuropathologies, and should also be of fundamental interest for investigations in other areas of systems biology.
Keywords: brain tumors; cellular interactions; cranial window; intravital imaging; multicolor transgenic mouse; spectral deconvolution; two-photon microscopy.
Figures
References
LinkOut - more resources
Full Text Sources
Other Literature Sources

