Integrated physical, genetic and genome map of chickpea (Cicer arietinum L.)
- PMID: 24610029
- PMCID: PMC4273598
- DOI: 10.1007/s10142-014-0363-6
Integrated physical, genetic and genome map of chickpea (Cicer arietinum L.)
Abstract
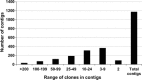
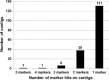
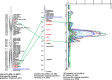
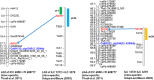
Physical map of chickpea was developed for the reference chickpea genotype (ICC 4958) using bacterial artificial chromosome (BAC) libraries targeting 71,094 clones (~12× coverage). High information content fingerprinting (HICF) of these clones gave high-quality fingerprinting data for 67,483 clones, and 1,174 contigs comprising 46,112 clones and 3,256 singletons were defined. In brief, 574 Mb genome size was assembled in 1,174 contigs with an average of 0.49 Mb per contig and 3,256 singletons represent 407 Mb genome. The physical map was linked with two genetic maps with the help of 245 BAC-end sequence (BES)-derived simple sequence repeat (SSR) markers. This allowed locating some of the BACs in the vicinity of some important quantitative trait loci (QTLs) for drought tolerance and reistance to Fusarium wilt and Ascochyta blight. In addition, fingerprinted contig (FPC) assembly was also integrated with the draft genome sequence of chickpea. As a result, ~965 BACs including 163 minimum tilling path (MTP) clones could be mapped on eight pseudo-molecules of chickpea forming 491 hypothetical contigs representing 54,013,992 bp (~54 Mb) of the draft genome. Comprehensive analysis of markers in abiotic and biotic stress tolerance QTL regions led to identification of 654, 306 and 23 genes in drought tolerance "QTL-hotspot" region, Ascochyta blight resistance QTL region and Fusarium wilt resistance QTL region, respectively. Integrated physical, genetic and genome map should provide a foundation for cloning and isolation of QTLs/genes for molecular dissection of traits as well as markers for molecular breeding for chickpea improvement.
Figures


Similar articles
-
A BAC/BIBAC-based physical map of chickpea, Cicer arietinum L.BMC Genomics. 2010 Sep 17;11:501. doi: 10.1186/1471-2164-11-501. BMC Genomics. 2010. PMID: 20849583 Free PMC article.
-
Genome-wide SNP discovery for development of high-density genetic map and QTL mapping of ascochyta blight resistance in chickpea (Cicer arietinum L.).Theor Appl Genet. 2019 Jun;132(6):1861-1872. doi: 10.1007/s00122-019-03322-3. Epub 2019 Mar 16. Theor Appl Genet. 2019. PMID: 30879097 Free PMC article.
-
mQTL-seq and classical mapping implicates the role of an AT-HOOK MOTIF CONTAINING NUCLEAR LOCALIZED (AHL) family gene in Ascochyta blight resistance of chickpea.Plant Cell Environ. 2018 Sep;41(9):2128-2140. doi: 10.1111/pce.13177. Epub 2018 May 16. Plant Cell Environ. 2018. PMID: 29492990
-
Advances in genetics and molecular breeding of three legume crops of semi-arid tropics using next-generation sequencing and high-throughput genotyping technologies.J Biosci. 2012 Nov;37(5):811-20. doi: 10.1007/s12038-012-9228-0. J Biosci. 2012. PMID: 23107917 Review.
-
Conventional and molecular breeding for disease resistance in chickpea: status and strategies.Biotechnol Genet Eng Rev. 2023 Oct;39(2):193-224. doi: 10.1080/02648725.2022.2110641. Epub 2022 Aug 12. Biotechnol Genet Eng Rev. 2023. PMID: 35959728 Review.
Cited by
-
Genetic Analysis of Partially Resistant and Susceptible Chickpea Cultivars in Response to Ascochyta rabiei Infection.Int J Mol Sci. 2024 Jan 22;25(2):1360. doi: 10.3390/ijms25021360. Int J Mol Sci. 2024. PMID: 38279360 Free PMC article.
-
Genotyping-by-sequencing based intra-specific genetic map refines a ''QTL-hotspot" region for drought tolerance in chickpea.Mol Genet Genomics. 2015 Apr;290(2):559-71. doi: 10.1007/s00438-014-0932-3. Epub 2014 Oct 25. Mol Genet Genomics. 2015. PMID: 25344290 Free PMC article.
-
MutMap Approach Enables Rapid Identification of Candidate Genes and Development of Markers Associated With Early Flowering and Enhanced Seed Size in Chickpea (Cicer arietinum L.).Front Plant Sci. 2021 Jul 12;12:688694. doi: 10.3389/fpls.2021.688694. eCollection 2021. Front Plant Sci. 2021. PMID: 34326857 Free PMC article.
-
A chickpea MAGIC population to dissect the genetics of complex traits.Plant Genome. 2025 Sep;18(3):e70096. doi: 10.1002/tpg2.70096. Plant Genome. 2025. PMID: 40851216 Free PMC article.
-
Genetic dissection of plant growth habit in chickpea.Funct Integr Genomics. 2017 Nov;17(6):711-723. doi: 10.1007/s10142-017-0566-8. Epub 2017 Jun 9. Funct Integr Genomics. 2017. PMID: 28600722
References
-
- Abu-Salem FM, Abou EA. Physico-chemical properties of tempeh produced from chickpea seeds. Arab J Am Sci. 2011;7:107–118.
-
- Anuradha C, Gaur PM, Pande S, Gali KK, Ganesh M, Kumar J, Varshney RK. Mapping QTL for resistance to Botrytis grey mould in chickpea. Euphytica. 2011;182:1–9. doi: 10.1007/s10681-011-0394-1. - DOI
-
- Arumuganathan K, Earle ED. Nuclear DNA content of some important plant species. Plant Mol Biol Report. 1991;9:208–218. doi: 10.1007/BF02672069. - DOI
-
- Aryamanesh N, Nelson MN, Yan G, Clarke HJ, Siddique KHM. Mapping a major gene for growth habit and QTLs for Ascochyta blight resistance and flowering time in a population between chickpea and Cicer reticulatum. Euphytica. 2010;173:307–319. doi: 10.1007/s10681-009-0086-2. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

