A comparison among the tissue-specific effects of aging and calorie restriction on TFAM amount and TFAM-binding activity to mtDNA in rat
- PMID: 24631828
- PMCID: PMC4335656
- DOI: 10.1016/j.bbagen.2014.03.004
A comparison among the tissue-specific effects of aging and calorie restriction on TFAM amount and TFAM-binding activity to mtDNA in rat
Abstract
Background: Mitochondrial Transcription Factor A (TFAM) is regarded as a histone-like protein of mitochondrial DNA (mtDNA), performing multiple functions for this genome. Aging affects mitochondria in a tissue-specific manner and only calorie restriction (CR) is able to delay or prevent the onset of several age-related changes also in mitochondria.
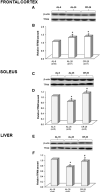
Methods: Samples of the frontal cortex and soleus skeletal muscle from 6- and 26-month-old ad libitum-fed and 26-month-old calorie-restricted rats and of the livers from 18- and 28-month-old ad libitum-fed and 28-month-old calorie-restricted rats were used to detect TFAM amount, TFAM-binding to mtDNA and mtDNA content.
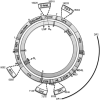
Results: We found an age-related increase in TFAM amount in the frontal cortex, not affected by CR, versus an age-related decrease in the soleus and liver, fully prevented by CR. The semi-quantitative analysis of in vivo binding of TFAM to specific mtDNA regions, by mtDNA immunoprecipitation assay and following PCR, showed a marked age-dependent decrease in TFAM-binding activity in the frontal cortex, partially prevented by CR. An age-related increase in TFAM-binding to mtDNA, fully prevented by CR, was found in the soleus and liver. MtDNA content presented a common age-related decrease, completely prevented by CR in the soleus and liver, but not in the frontal cortex.
Conclusions: The modulation of TFAM expression, TFAM-binding to mtDNA and mtDNA content with aging and CR showed a trend shared by the skeletal muscle and liver, but not by the frontal cortex counterpart.
General significance: Aging and CR appear to induce similar mitochondrial molecular mechanisms in the skeletal muscle and liver, different from those elicited in the frontal cortex.
Keywords: Aging rat; Calorie restriction; Mitochondrial Transcription Factor A; Mitochondrial Transcription Factor A–mitochondrial deoxyribonucleic acid binding; Tissue-specificity.
Copyright © 2014 Elsevier B.V. All rights reserved.
Figures


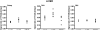

References
-
- Larsson NG, Garman JD, Oldfors A, Barsh GS, Clayton DA. A single mouse gene encodes the mitochondrial transcription factor A and a testis-specific nuclear HMG-box protein. Nat. Genet. 1996;13:296–302. - PubMed
-
- Lezza AMS. Mitochondrial transcription factor A (TFAM): one actor for different roles. Front. Biol. 2012;7:30–39.
-
- Parisi MA, Clayton DA. Similarity of human mitochondrial transcription factor 1 to high mobility group proteins. Science. 1991;252:965–969. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

