High precision prediction of functional sites in protein structures
- PMID: 24632601
- PMCID: PMC3954699
- DOI: 10.1371/journal.pone.0091240
High precision prediction of functional sites in protein structures
Abstract
We address the problem of assigning biological function to solved protein structures. Computational tools play a critical role in identifying potential active sites and informing screening decisions for further lab analysis. A critical parameter in the practical application of computational methods is the precision, or positive predictive value. Precision measures the level of confidence the user should have in a particular computed functional assignment. Low precision annotations lead to futile laboratory investigations and waste scarce research resources. In this paper we describe an advanced version of the protein function annotation system FEATURE, which achieved 99% precision and average recall of 95% across 20 representative functional sites. The system uses a Support Vector Machine classifier operating on the microenvironment of physicochemical features around an amino acid. We also compared performance of our method with state-of-the-art sequence-level annotator Pfam in terms of precision, recall and localization. To our knowledge, no other functional site annotator has been rigorously evaluated against these key criteria. The software and predictive models are incorporated into the WebFEATURE service at http://feature.stanford.edu/wf4.0-beta.
Conflict of interest statement
Figures
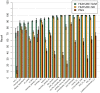
References
-
- Thornton J (2001) Structural genomics takes off. Trends Biochem Sci 26: 88–89. - PubMed
-
- Fetrow JS, Skolnick J (1998) Method for prediction of protein function from sequence using the sequence-to-structure-to-function paradigm with application to glutaredoxins/thioredoxins and T1 ribonucleases. J Mol Biol 281: 949–968. - PubMed
-
- Polacco BJ, Babbitt PC (2006) Automated discovery of 3D motifs for protein function annotation. Bioinformatics 22: 723–730. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

