Optimized three-dimensional sodium imaging of the human heart on a clinical 3T scanner
- PMID: 24639022
- PMCID: PMC4167174
- DOI: 10.1002/mrm.25175
Optimized three-dimensional sodium imaging of the human heart on a clinical 3T scanner
Abstract
Purpose: Optimization of sequence and sequence parameters to allow three-dimensional (3D) sodium imaging of the entire human heart in vivo in a clinically reasonable time.
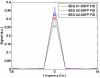
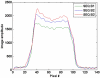
Theory and methods: A stack of spirals pulse sequence was optimized for cardiac imaging by considering factors such as spoiling, nutation angles, repetition time, echo time, T1/T2 relaxation, off-resonance, data acquisition window, motion, and segmented k-space acquisition. Simulations based on Bloch equations as well as the exact trajectory used for data acquisition provided the basis for choice of parameter combinations for sodium imaging. Sodium phantom scanning was used to validate the choice of parameters and for corroboration with simulations. In vivo cardiac imaging in six volunteers was also performed with an optimized sequence.
Results: Phantom studies showed good correlation with simulation results. Images obtained from human volunteers showed that the heart can be imaged with a nominal resolution of 5 × 5 × 10 mm(3) and with a signal-to-noise ratio >15 (in the septum) in about 6-10 minutes. Long axis views of the reformatted human heart show true 3D imaging capability.
Conclusion: Optimization of the sequence and its parameters allowed in vivo 3D sodium imaging of the entire human heart in a clinically reasonable time.
Keywords: 3D imaging; heart; sodium MRI.
© 2014 Wiley Periodicals, Inc.
Figures










References
-
- Parrish TB, Fieno DS, Fitzgerald SW, Judd RM. Theoretical basis for sodium and potassium MRI of the human heart at 1.5 T. Magn Reson Med. 1997;38(4):653–661. - PubMed
-
- Polimeni PI. Extracellular space and ionic distribution in rat ventricle. Am J Physiol. 1974;227(3):676–683. - PubMed
-
- Ouwerkerk R. Sodium MRI. Methods Mol Biol. 2011;711:175–201. - PubMed
-
- Rochitte CE, Kim RJ, Hillenbrand HB, Chen EL, Lima JA. Microvascular integrity and the time course of myocardial sodium accumulation after acute infarction. Circ Res. 2000;87(8):648–655. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

