Signaling networks in palate development
- PMID: 24644145
- PMCID: PMC3991764
- DOI: 10.1002/wsbm.1265
Signaling networks in palate development
Abstract
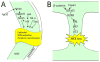
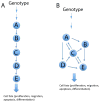
Palatogenesis, the formation of the palate, is a dynamic process regulated by a complex series of context-dependent morphogenetic signaling events. Many genes involved in palatogenesis have been discovered through the use of genetically manipulated mouse models as well as from human genetic studies, but the roles of these genes and their products in signaling networks regulating palatogenesis are still poorly known. In this review, we give a brief overview on palatogenesis and introduce key signaling cascades leading to formation of the intact palate. Moreover, we review conceptual differences between pathway biology and network biology and discuss how some of the recent technological advances in conjunction with mouse genetic models have contributed to our understanding of signaling networks regulating palate growth and fusion. For further resources related to this article, please visit the WIREs website.
Conflict of interest: The authors have declared no conflicts of interest for this article.
© 2014 Wiley Periodicals, Inc.
Figures


References
-
- Murray JC. Gene/environment causes of cleft lip and/or palate. Clin Genet. 2002;61:248–256. - PubMed
-
- Schutte BC, Murray JC. The many faces and factors of orofacial clefts. Hum Mol Genet. 1999;8:1853–1859. - PubMed
-
- Gritli-Linde A. Molecular control of secondary palate development. Dev Biol. 2007;301:309–326. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

