Unique features of the loblolly pine (Pinus taeda L.) megagenome revealed through sequence annotation
- PMID: 24653211
- PMCID: PMC3948814
- DOI: 10.1534/genetics.113.159996
Unique features of the loblolly pine (Pinus taeda L.) megagenome revealed through sequence annotation
Abstract
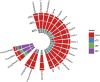
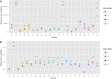
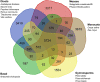
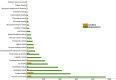
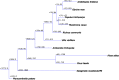
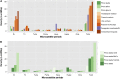
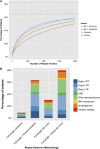
The largest genus in the conifer family Pinaceae is Pinus, with over 100 species. The size and complexity of their genomes (∼20-40 Gb, 2n = 24) have delayed the arrival of a well-annotated reference sequence. In this study, we present the annotation of the first whole-genome shotgun assembly of loblolly pine (Pinus taeda L.), which comprises 20.1 Gb of sequence. The MAKER-P annotation pipeline combined evidence-based alignments and ab initio predictions to generate 50,172 gene models, of which 15,653 are classified as high confidence. Clustering these gene models with 13 other plant species resulted in 20,646 gene families, of which 1554 are predicted to be unique to conifers. Among the conifer gene families, 159 are composed exclusively of loblolly pine members. The gene models for loblolly pine have the highest median and mean intron lengths of 24 fully sequenced plant genomes. Conifer genomes are full of repetitive DNA, with the most significant contributions from long-terminal-repeat retrotransposons. In depth analysis of the tandem and interspersed repetitive content yielded a combined estimate of 82%.
Keywords: conifer; gene family; introns; repeats; retrotransposons.
Figures
Similar articles
-
Decoding the massive genome of loblolly pine using haploid DNA and novel assembly strategies.Genome Biol. 2014 Mar 4;15(3):R59. doi: 10.1186/gb-2014-15-3-r59. Genome Biol. 2014. PMID: 24647006 Free PMC article.
-
The Pinus taeda genome is characterized by diverse and highly diverged repetitive sequences.BMC Genomics. 2010 Jul 7;11:420. doi: 10.1186/1471-2164-11-420. BMC Genomics. 2010. PMID: 20609256 Free PMC article.
-
Insights into the loblolly pine genome: characterization of BAC and fosmid sequences.PLoS One. 2013 Sep 4;8(9):e72439. doi: 10.1371/journal.pone.0072439. eCollection 2013. PLoS One. 2013. PMID: 24023741 Free PMC article.
-
Insights into conifer giga-genomes.Plant Physiol. 2014 Dec;166(4):1724-32. doi: 10.1104/pp.114.248708. Epub 2014 Oct 27. Plant Physiol. 2014. PMID: 25349325 Free PMC article. Review.
-
Genome histories clarify evolution of the expansin superfamily: new insights from the poplar genome and pine ESTs.J Plant Res. 2006 Jan;119(1):11-21. doi: 10.1007/s10265-005-0253-z. Epub 2006 Jan 13. J Plant Res. 2006. PMID: 16411016 Review.
Cited by
-
Analysis of the giant genomes of Fritillaria (Liliaceae) indicates that a lack of DNA removal characterizes extreme expansions in genome size.New Phytol. 2015 Oct;208(2):596-607. doi: 10.1111/nph.13471. Epub 2015 Jun 8. New Phytol. 2015. PMID: 26061193 Free PMC article.
-
Comprehensive collection of genes and comparative analysis of full-length transcriptome sequences from Japanese larch (Larix kaempferi) and Kuril larch (Larix gmelinii var. japonica).BMC Plant Biol. 2022 Oct 4;22(1):470. doi: 10.1186/s12870-022-03862-9. BMC Plant Biol. 2022. PMID: 36192701 Free PMC article.
-
Landscape of Fluid Sets of Hairpin-Derived 21-/24-nt-Long Small RNAs at Seed Set Uncovers Special Epigenetic Features in Picea glauca.Genome Biol Evol. 2017 Jan 1;9(1):82-92. doi: 10.1093/gbe/evw283. Genome Biol Evol. 2017. PMID: 28082604 Free PMC article.
-
Next Generation Sequencing Technologies: The Doorway to the Unexplored Genomics of Non-Model Plants.Front Plant Sci. 2015 Dec 16;6:1074. doi: 10.3389/fpls.2015.01074. eCollection 2015. Front Plant Sci. 2015. PMID: 26734016 Free PMC article. Review.
-
Application of crop wild relatives in modern breeding: An overview of resources, experimental and computational methodologies.Front Plant Sci. 2022 Nov 17;13:1008904. doi: 10.3389/fpls.2022.1008904. eCollection 2022. Front Plant Sci. 2022. PMID: 36466237 Free PMC article. Review.
References
-
- Ahuja M. R., Neale D. B., 2005. Evolution of genome size in conifers. Silvae Genet. 54: 126–137.
-
- Altschul S. F., Gish W., Miller W., Myers E. W., Lipman D. J., 1990. Basic local alignment search tool. J. Mol. Biol. 215: 403–410. - PubMed
-
- Aronen T., Ryynanen L., 2012. Variation in telomeric repeats of Scots pine (Pinus sylvestris L.). Tree Genet. Genomes 8: 267–275.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

