Comparative population genomics: power and principles for the inference of functionality
- PMID: 24656563
- PMCID: PMC4073783
- DOI: 10.1016/j.tig.2014.02.002
Comparative population genomics: power and principles for the inference of functionality
Abstract
The availability of sequenced genomes from multiple related organisms allows the detection and localization of functional genomic elements based on the idea that such elements evolve more slowly than neutral sequences. Although such comparative genomics methods have proven useful in discovering functional elements and ascertaining levels of functional constraint in the genome as a whole, here we outline limitations intrinsic to this approach that cannot be overcome by sequencing more species. We argue that it is essential to supplement comparative genomics with ultra-deep sampling of populations from closely related species to enable substantially more powerful genomic scans for functional elements. The convergence of sequencing technology and population genetics theory has made such projects feasible and has exciting implications for functional genomics.
Copyright © 2014 Elsevier Ltd. All rights reserved.
Figures
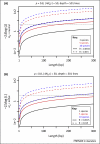
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

