Internal translation initiation and eIF4F/ATP-independent scanning of mRNA by eukaryotic ribosomal particles
- PMID: 24657959
- PMCID: PMC3963034
- DOI: 10.1038/srep04438
Internal translation initiation and eIF4F/ATP-independent scanning of mRNA by eukaryotic ribosomal particles
Abstract
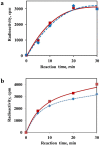
The recombinant mRNAs with 5'-untranslated region, called omega leader, of tobacco mosaic virus RNA are known to be well translated in eukaryotic cell-free systems, even if deprived of cap structure. Using the method of primer extension inhibition (toe-printing), the ribosomal particles were shown to initiate translation at uncapped omega leader when its 5'-end was blocked by a stable RNA-DNA double helix, thus providing evidence for internal initiation. The scanning of the leader sequence and the formation of ribosomal 48S initiation complexes at the initiation AUG codon occurred in the absence of ATP-dependent initiation factor eIF4F, as well as without ATP. The latter results implied the ability of ribosomal initiation complexes for ATP-independent, diffusional wandering (also called bi-directional movement) along the leader sequence during scanning.
Figures




Similar articles
-
Migration of Small Ribosomal Subunits on the 5' Untranslated Regions of Capped Messenger RNA.Int J Mol Sci. 2019 Sep 10;20(18):4464. doi: 10.3390/ijms20184464. Int J Mol Sci. 2019. PMID: 31510048 Free PMC article.
-
[Modification of the 5' End of mRNA Leader Sequence Alters the Set of Initiation Factors Essential for Initiation of Translation].Mol Biol (Mosk). 2020 May-Jun;54(3):480-486. doi: 10.31857/S0026898420030143. Mol Biol (Mosk). 2020. PMID: 32492012 Russian.
-
The roles of individual eukaryotic translation initiation factors in ribosomal scanning and initiation codon selection.Genes Dev. 2002 Nov 15;16(22):2906-22. doi: 10.1101/gad.1020902. Genes Dev. 2002. PMID: 12435632 Free PMC article.
-
A Cap for Every Occasion: Alternative eIF4F Complexes.Trends Biochem Sci. 2016 Oct;41(10):821-823. doi: 10.1016/j.tibs.2016.05.009. Epub 2016 Jun 6. Trends Biochem Sci. 2016. PMID: 27283511 Free PMC article. Review.
-
Structural Insights into the Mechanism of Scanning and Start Codon Recognition in Eukaryotic Translation Initiation.Trends Biochem Sci. 2017 Aug;42(8):589-611. doi: 10.1016/j.tibs.2017.03.004. Epub 2017 Apr 22. Trends Biochem Sci. 2017. PMID: 28442192 Review.
Cited by
-
RNAi-Based Biocontrol of Wheat Nematodes Using Natural Poly-Component Biostimulants.Front Plant Sci. 2019 Apr 17;10:483. doi: 10.3389/fpls.2019.00483. eCollection 2019. Front Plant Sci. 2019. PMID: 31057585 Free PMC article.
-
Yeast eIF4A enhances recruitment of mRNAs regardless of their structural complexity.Elife. 2017 Nov 30;6:e31476. doi: 10.7554/eLife.31476. Elife. 2017. PMID: 29192585 Free PMC article.
-
Enhanced or reversible RNA N6-methyladenosine editing by red/far-red light induction.Nucleic Acids Res. 2025 Feb 27;53(5):gkaf181. doi: 10.1093/nar/gkaf181. Nucleic Acids Res. 2025. PMID: 40103228 Free PMC article.
-
Making ends meet: New functions of mRNA secondary structure.Wiley Interdiscip Rev RNA. 2021 Mar;12(2):e1611. doi: 10.1002/wrna.1611. Epub 2020 Jun 29. Wiley Interdiscip Rev RNA. 2021. PMID: 32597020 Free PMC article. Review.
-
Inhibition of Cell-Free Translation and Replication of Tobacco Mosaic Virus RNA by Exogenously Added 5'-Proximal Fragments of the Genomic RNA.Viruses. 2022 Sep 4;14(9):1962. doi: 10.3390/v14091962. Viruses. 2022. PMID: 36146772 Free PMC article.
References
-
- Pestova T. V., Lorsch J. R. & Hellen C. U. T. [The mechanism of translation initiation in eukaryotes]. Translational Control in Biology and Medicine [Mathews, M. B., Sonenberg, N. & Hershey, J. W. B. (eds)] [87–128] (Cold Spring Harbor Lab Press, Cold Spring Harbor NY, 2007).
-
- Kozak M. How do eucaryotic ribosomes select initiation regions in messenger RNA? Cell 15, 1109–1123 (1978). - PubMed
-
- Kozak M. Role of ATP in binding and migration of 40S ribosomal subunits. Cell 22, 459–467 (1980). - PubMed
-
- Doudna J. A. & Sarnow P. [Translation initiation by viral internal ribosome entry sites]. Translational Control in Biology and Medicine, [Mathews, M. B., Sonenberg, N. & Hershey, J. W. B. (eds)] [129–153] (Cold Spring Harbor Lab Press, Cold Spring Harbor NY) (2007).
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

