Multifactorial comparative proteomic study of cytochrome P450 2E1 function in chronic alcohol administration
- PMID: 24658151
- PMCID: PMC3962406
- DOI: 10.1371/journal.pone.0092504
Multifactorial comparative proteomic study of cytochrome P450 2E1 function in chronic alcohol administration
Abstract
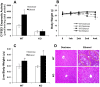
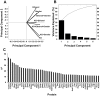
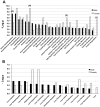
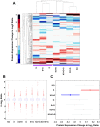
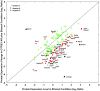
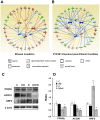
With the use of iTRAQ technique, a multifactorial comparative proteomic study can be performed. In this study, to obtain an overview of ethanol, CYP2E1 and gender effects on liver injury and gain more insight into the underlying molecular mechanism, mouse liver proteomes were quantitatively analyzed using iTRAQ under eight conditions including mice of different genders, wild type versus CYP2E1 knockout, and normal versus alcohol diet. A series of statistical and bioinformatic analyses were explored to simplify and clarify multifactorial comparative proteomic data. First, with the Principle Component analysis, six proteins, CYP2E1, FAM25, CA3, BHMT, HIBADH and ECHS1, involved in oxidation reduction, energy and lipid metabolism and amino acid metabolism, were identified as the most differentially expressed gene products across all of the experimental conditions of our chronic alcoholism model. Second, hierarchical clustering analysis showed CYP2E1 knockout played a primary role in the overall differential protein expression compared with ethanol and gender factors. Furthermore, pair-wise multiple comparisons have revealed that the only significant expression difference lied in wild-type and CYP2E1 knockout mice both treated with ethanol. Third, K-mean clustering analysis indicated that the CYP2E1 knockout had the reverse effect on ethanol induced oxidative stress and lipid oxidation. More importantly, IPA analysis of proteomic data inferred that the gene expressions of two upstream regulators, NRF2 and PPARα, regulated by chronic alcohol feeding and CYP2E1 knockout, are involved in ethanol induced oxidative stress and lipid oxidation. The present study provides an effectively comprehensive data analysis strategy to compare multiple biological factors, contributing to biochemical effects of alcohol on the liver. The mass spectrometry proteomics data have been deposited to the ProteomeXchange with data set identifier of PXD000635.
Conflict of interest statement
Figures

References
-
- Lieber CS (1984) Metabolism and metabolic effects of alcohol. Med Clin North Am 68: 3–31. - PubMed
-
- Ramaiah S, Rivera C, Arteel G (2004) Early-phase alcoholic liver disease: an update on animal models, pathology, and pathogenesis. Int J Toxicol 23: 217–231. - PubMed
-
- Nanji AA, Su GL, Laposata M, French SW (2002) Pathogenesis of alcoholic liver disease—recent advances. Alcohol Clin Exp Res 26: 731–736. - PubMed
-
- Lieber CS (2004) Alcoholic fatty liver: its pathogenesis and mechanism of progression to inflammation and fibrosis. Alcohol 34: 9–19. - PubMed
-
- Diehl AM (2002) Liver disease in alcohol abusers: clinical perspective. Alcohol 27: 7–11. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

