Discovering new bioactive molecules from microbial sources
- PMID: 24661414
- PMCID: PMC3992017
- DOI: 10.1111/1751-7915.12123
Discovering new bioactive molecules from microbial sources
Abstract
There is an increased need for new drug leads to treat diseases in humans, animals and plants. A dramatic example is represented by the need for novel and more effective antibiotics to combat multidrug-resistant microbial pathogens. Natural products represent a major source of approved drugs and still play an important role in supplying chemical diversity, despite a decreased interest by large pharmaceutical companies. Novel approaches must be implemented to decrease the chances of rediscovering the tens of thousands of known natural products. In this review, we present an overview of natural product screening, focusing particularly on microbial products. Different approaches can be implemented to increase the probability of finding new bioactive molecules. We thus present the rationale and selected examples of the use of hypersensitive assays; of accessing unexplored microorganisms, including the metagenome; and of genome mining. We then focus our attention on the technology platform that we are currently using, consisting of approximately 70,000 microbial strains, mostly actinomycetes and filamentous fungi, and discuss about high-quality screening in the search for bioactive molecules. Finally, two case studies are discussed, including the spark that arose interest in the compound: in the case of orthoformimycin, the novel mechanism of action predicted a novel structural class; in the case of NAI-112, structural similarity pointed out to a possible in vivo activity. Both predictions were then experimentally confirmed.
© 2014 The Authors. Microbial Biotechnology published by John Wiley & Sons Ltd and Society for Applied Microbiology.
Figures

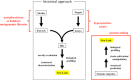

Percent distribution of the approximately 70 000 strains.
High-level distribution of the approximately 3000 classified filamentous fungi.
Suborder-level distribution of the approximately 43 000 actinomycetes.

Fluorescence stopped-flow kinetics of the mRNA and
Phe-tRNA EF-G-dependent translocation in the absence (black trace) or in the presence of 20 μM of orthoformimycin (red trace).
Structure of orthoformimycin.
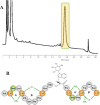
HPLC trace of the fractionated extract from Actinoplanes ID112781. The only peak with antibacterial activity is highlighted.
Chemical structure of NAI-112. The labionine (Lab) rings AB and A′B′ are highlighted, together with dehydrated amino acids Dha (dehydroalanine) and Dhb (dehydrobutyrine). The N-glycosylated Trp residue is detailed.
References
-
- Aigle B, Corre C. Waking up Streptomyces secondary metabolism by constitutive expression of activators or genetic disruption of repressors. Methods Enzymol. 2012;517:343–366. - PubMed
-
- Baltz RH. Antibiotic discovery from actinomycetes: will a renaissance follow the decline and fall? SIM News. 2005;55:186–196.
-
- Baltz RH. Antimicrobials from actinomycetes: back to the future. Microbe. 2007;2:125–131.
-
- Baltz RH. Renaissance in antibacterial discovery from actinomycetes. Curr Opin Pharmacol. 2008;8:557–563. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

