Genetic host specificity of hepatitis E virus
- PMID: 24667049
- PMCID: PMC5745802
- DOI: 10.1016/j.meegid.2014.03.011
Genetic host specificity of hepatitis E virus
Abstract
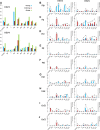
Hepatitis E virus (HEV) causes epidemic and sporadic cases of hepatitis worldwide. HEV genotypes 3 (HEV3) and 4 (HEV4) infect humans and animals, with swine being the primary reservoir. The relevance of HEV genetic diversity to host adaptation is poorly understood. We employed a Bayesian network (BN) analysis of HEV3 and HEV4 to detect epistatic connectivity among protein sites and its association with the host specificity in each genotype. The data imply coevolution among ∼70% of polymorphic sites from all HEV proteins and association of numerous coevolving sites with adaptation to swine or humans. BN models for individual proteins and domains of the nonstructural polyprotein detected the host origin of HEV strains with accuracy of 74-93% and 63-87%, respectively. These findings, taken together with lack of phylogenetic association to host, suggest that the HEV host specificity is a heritable and convergent phenotypic trait achievable through variety of genetic pathways (abundance), and explain a broad host range for HEV3 and HEV4.
Keywords: Adaptation; Bayesian network; Coevolution; HEV ORFs; Hepatitis E virus; Prediction.
Published by Elsevier B.V.
Figures



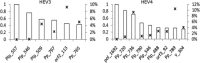


Similar articles
-
Swine hepatitis E virus: Cross-species infection, pork safety and chronic infection.Virus Res. 2020 Jul 15;284:197985. doi: 10.1016/j.virusres.2020.197985. Epub 2020 Apr 23. Virus Res. 2020. PMID: 32333941 Free PMC article. Review.
-
Analysing complete genome sequence of swine hepatitis E virus (HEV), strain CHN-XJ-SW13 isolated from Xinjiang, China: putative host range, and disease severity determinants in HEV.Infect Genet Evol. 2011 Apr;11(3):618-23. doi: 10.1016/j.meegid.2011.01.018. Epub 2011 Feb 1. Infect Genet Evol. 2011. PMID: 21292034
-
Genotype-Specific Evolution of Hepatitis E Virus.J Virol. 2017 Apr 13;91(9):e02241-16. doi: 10.1128/JVI.02241-16. Print 2017 May 1. J Virol. 2017. PMID: 28202767 Free PMC article.
-
Identity of a novel swine hepatitis E virus in Taiwan forming a monophyletic group with Taiwan isolates of human hepatitis E virus.J Clin Microbiol. 1999 Dec;37(12):3828-34. doi: 10.1128/JCM.37.12.3828-3834.1999. J Clin Microbiol. 1999. PMID: 10565892 Free PMC article.
-
Hepatitis E Virus Genotypes and Evolution: Emergence of Camel Hepatitis E Variants.Int J Mol Sci. 2017 Apr 20;18(4):869. doi: 10.3390/ijms18040869. Int J Mol Sci. 2017. PMID: 28425927 Free PMC article. Review.
Cited by
-
Zoonotic Hepatitis E Virus: Classification, Animal Reservoirs and Transmission Routes.Viruses. 2016 Oct 3;8(10):270. doi: 10.3390/v8100270. Viruses. 2016. PMID: 27706110 Free PMC article. Review.
-
A Novel Hepe-Like Virus from Farmed Giant Freshwater Prawn Macrobrachium rosenbergii.Viruses. 2020 Mar 17;12(3):323. doi: 10.3390/v12030323. Viruses. 2020. PMID: 32192159 Free PMC article.
-
Genetic Variability and Evolution of Hepatitis E Virus.Viruses. 2019 May 18;11(5):456. doi: 10.3390/v11050456. Viruses. 2019. PMID: 31109076 Free PMC article. Review.
-
Quantitative differences between intra-host HCV populations from persons with recently established and persistent infections.Virus Evol. 2020 Dec 30;7(1):veaa103. doi: 10.1093/ve/veaa103. eCollection 2021 Jan. Virus Evol. 2020. PMID: 33505710 Free PMC article.
-
Swine hepatitis E virus: Cross-species infection, pork safety and chronic infection.Virus Res. 2020 Jul 15;284:197985. doi: 10.1016/j.virusres.2020.197985. Epub 2020 Apr 23. Virus Res. 2020. PMID: 32333941 Free PMC article. Review.
References
-
- Aggarwal R. Clinical presentation of hepatitis E. Virus Res. 2011;161:15–22. - PubMed
-
- Aggarwal R., Kamili S., Spelbring J., Krawczynski K. Experimental studies on subclinical hepatitis E virus infection in cynomolgus macaques. J. Infect. Dis. 2001;184:1380–1385. - PubMed
-
- Arankalle V.A., Chobe L.P., Joshi M.V., Chadha M.S., Kundu B., Walimbe A.M. Human and swine hepatitis E viruses from Western India belong to different genotypes. J. Hepatol. 2002;36:417–425. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

