Test of rare variant association based on affected sib-pairs
- PMID: 24667785
- PMCID: PMC4297896
- DOI: 10.1038/ejhg.2014.43
Test of rare variant association based on affected sib-pairs
Abstract
With the development of sequencing techniques, there is increasing interest to detect associations between rare variants and complex traits. Quite a few statistical methods to detect associations between rare variants and complex traits have been developed for unrelated individuals. Statistical methods for detecting rare variant associations under family-based designs have not received as much attention as methods for unrelated individuals. Recent studies show that rare disease variants will be enriched in family data and thus family-based designs may improve power to detect rare variant associations. In this article, we propose a novel test to test association between the optimally weighted combination of variants and trait of interests for affected sib-pairs. The optimal weights are analytically derived and can be calculated from sampled genotypes and phenotypes. Based on the optimal weights, the proposed method is robust to the directions of the effects of causal variants and is less affected by neutral variants than existing methods are. Our simulation results show that, in all the cases, the proposed method is substantially more powerful than existing methods based on unrelated individuals and existing methods based on affected sib-pairs.
Figures
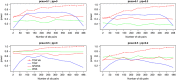
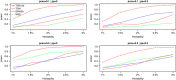
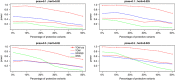
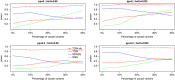
Similar articles
-
Detecting association of rare variants by testing an optimally weighted combination of variants for quantitative traits in general families.Ann Hum Genet. 2013 Nov;77(6):524-34. doi: 10.1111/ahg.12038. Epub 2013 Aug 22. Ann Hum Genet. 2013. PMID: 23968488 Free PMC article.
-
A modified association test for rare and common variants based on affected sib-pair design.J Theor Biol. 2019 Apr 21;467:1-6. doi: 10.1016/j.jtbi.2019.01.014. Epub 2019 Jan 29. J Theor Biol. 2019. PMID: 30707975
-
Two adaptive weighting methods to test for rare variant associations in family-based designs.Genet Epidemiol. 2012 Jul;36(5):499-507. doi: 10.1002/gepi.21646. Epub 2012 Jun 1. Genet Epidemiol. 2012. PMID: 22674630
-
Identifying rare variants associated with complex traits via sequencing.Curr Protoc Hum Genet. 2013 Jul;Chapter 1:Unit 1.26. doi: 10.1002/0471142905.hg0126s78. Curr Protoc Hum Genet. 2013. PMID: 23853079 Free PMC article. Review.
-
Two-phase and family-based designs for next-generation sequencing studies.Front Genet. 2013 Dec 13;4:276. doi: 10.3389/fgene.2013.00276. Front Genet. 2013. PMID: 24379824 Free PMC article. Review.
Cited by
-
Gene-based and pathway-based testing for rare-variant association in affected sib pairs.Genet Epidemiol. 2020 Jun;44(4):368-381. doi: 10.1002/gepi.22291. Epub 2020 Apr 1. Genet Epidemiol. 2020. PMID: 32237178 Free PMC article.
-
Association detection between multiple traits and rare variants based on family data via a nonparametric method.PeerJ. 2023 Sep 26;11:e16040. doi: 10.7717/peerj.16040. eCollection 2023. PeerJ. 2023. PMID: 37780393 Free PMC article.
References
-
- McCarthy MI, Abecasis GR, Cardon LR, et al. Genome-wide association studies for complex traits: consensus, uncertainty and challenges. Nat Rev Genet. 2008;9:356–369. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

