Redefinition of the human mast cell transcriptome by deep-CAGE sequencing
- PMID: 24671954
- PMCID: PMC3999759
- DOI: 10.1182/blood-2013-02-483792
Redefinition of the human mast cell transcriptome by deep-CAGE sequencing
Abstract
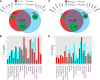
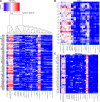
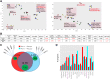
Mast cells (MCs) mature exclusively in peripheral tissues, hampering research into their developmental and functional programs. Here, we employed deep cap analysis of gene expression on skin-derived MCs to generate the most comprehensive view of the human MC transcriptome ever reported. An advantage is that MCs were embedded in the FANTOM5 project, giving the opportunity to contrast their molecular signature against a multitude of human samples. We demonstrate that MCs possess a unique and surprising transcriptional landscape, combining hematopoietic genes with those exclusively active in MCs and genes not previously reported as expressed by MCs (several of them markers of unrelated tissues). We also found functional bone morphogenetic protein receptors transducing activatory signals in MCs. Conversely, several immune-related genes frequently studied in MCs were not expressed or were weakly expressed. Comparing MCs ex vivo with cultured counterparts revealed profound changes in the MC transcriptome in in vitro surroundings. We also determined the promoter usage of MC-expressed genes and identified associated motifs active in the lineage. Befitting their uniqueness, MCs had no close relative in the hematopoietic network (also only distantly related with basophils). This rich data set reveals that our knowledge of human MCs is still limited, but with this resource, novel functional programs of MCs may soon be discovered.
Figures

Comment in
-
Deep sequencing blood transcriptomes.Blood. 2014 Apr 24;123(17):2595-6. doi: 10.1182/blood-2014-03-563452. Blood. 2014. PMID: 24764555 No abstract available.
References
-
- Brown JM, Wilson TM, Metcalfe DD. The mast cell and allergic diseases: role in pathogenesis and implications for therapy. Clin Exp Allergy. 2008;38(1):4–18. - PubMed
-
- Navi D, Saegusa J, Liu FT. Mast cells and immunological skin diseases. Clin Rev Allergy Immunol. 2007;33(1-2):144–155. - PubMed
-
- Sayed BA, Christy A, Quirion MR, Brown MA. The master switch: the role of mast cells in autoimmunity and tolerance. Annu Rev Immunol. 2008;26:705–739. - PubMed
-
- Kneilling M, Röcken M. Mast cells: novel clinical perspectives from recent insights. Exp Dermatol. 2009;18(5):488–496. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

