Metabolic gene profile in early human fetal heart development
- PMID: 24674993
- PMCID: PMC11514182
- DOI: 10.1093/molehr/gau026
Metabolic gene profile in early human fetal heart development
Abstract
The primitive cardiac tube starts beating 6-8 weeks post fertilization in the developing embryo. In order to describe normal cardiac development during late first and early second trimester in human fetuses this study used microarray and pathways analysis and created a corresponding 'normal' database. Fourteen fetal hearts from human fetuses between 10 and 18 weeks of gestational age (GA) were prospectively collected at the time of elective termination of pregnancy. RNA from recovered tissues was used for transcriptome analysis with Affymetrix 1.0 ST microarray chip. From the amassed data we investigated differences in cardiac development within the 10-18 GA period dividing the sample by GA in three groups: 10-12 (H1), 13-15 (H2) and 16-18 (H3) weeks. A fold change of 2 or above adjusted for a false discovery rate of 5% was used as initial cutoff to determine differential gene expression for individual genes. Test for enrichment to identify functional groups was carried out using the Gene Ontology (GO) and the Kyoto Encyclopedia of Genes and Genomes (KEGG). Array analysis correctly identified the cardiac specific genes, and transcripts reported to be differentially expressed were confirmed by qRT-PCR. Single transcript and Ontology analysis showed first trimester heart expression of myosin-related genes to be up-regulated >5-fold compared with second trimester heart. In contrast the second trimester hearts showed further gestation-related increases in many genes involved in energy production and cardiac remodeling. In conclusion, fetal heart development during the first trimester was dominated by heart-specific genes coding for myocardial development and differentiation. During the second trimester, transcripts related to energy generation and cardiomyocyte communication for contractile coordination/proliferation were more dominant. Transcripts related to fatty acid metabolism can be seen as early as 10 weeks and clearly increase as the heart matures. Retinol receptor and gamma-aminobutyric acid (GABA) receptor transcripts were detected, and have not been described previously in human fetal heart during this period. For the first time global gene expression of heart has been described in human samples to create a database of normal development to understand and compare with known abnormal fetal heart development.
Keywords: cardiac development; fetal; human; microarray; renin-angiotensin.
© The Author 2014. Published by Oxford University Press on behalf of the European Society of Human Reproduction and Embryology. All rights reserved. For Permissions, please email: journals.permissions@oup.com.
Conflict of interest statement
None declared.
Figures
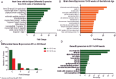


References
-
- Gratacos E Gomez R Nicolaides K Romero R Cabero L. Medicina Fetal. Spain: Editorial Medica Panamericana; 2007. 822.
-
- Huber W Gentleman R matchprobes: a Bioconductor package for the sequence-matching of microarray probe elements. Bioinformatics. 2004;20:1651–1652. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

