Non-cell-autonomous regulation of root hair patterning genes by WRKY75 in Arabidopsis
- PMID: 24676857
- PMCID: PMC4012579
- DOI: 10.1104/pp.113.233775
Non-cell-autonomous regulation of root hair patterning genes by WRKY75 in Arabidopsis
Abstract
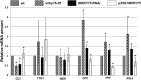
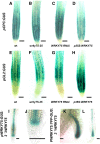
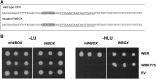
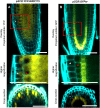
In Arabidopsis (Arabidopsis thaliana), root hairs are formed in cell files over the cleft of underlying cortex cells. This pattern is established by a well-known gene regulatory network of transcription factors. In this study, we show that WRKY75 suppresses root hair development in nonroot hair files and that it represses the expression of TRIPTYCHON and CAPRICE. The WRKY75 protein binds to the CAPRICE promoter in a yeast one-hybrid assay. Binding to the promoter fragment requires an intact WRKY protein-binding motif, the W box. A comparison of the spatial expression of WRKY75 and the localization of the WRKY75 protein revealed that WRKY75 is expressed in the pericycle and vascular tissue and that the WRKY75 RNA or protein moves into the epidermis.
Figures



References
-
- Bernhardt C, Lee MM, Gonzalez A, Zhang F, Lloyd A, Schiefelbein J. (2003) The bHLH genes GLABRA3 (GL3) and ENHANCER OF GLABRA3 (EGL3) specify epidermal cell fate in the Arabidopsis root. Development 130: 6431–6439 - PubMed
-
- Bernhardt C, Zhao M, Gonzalez A, Lloyd A, Schiefelbein J. (2005) The bHLH genes GL3 and EGL3 participate in an intercellular regulatory circuit that controls cell patterning in the Arabidopsis root epidermis. Development 132: 291–298 - PubMed
-
- Birnbaum K, Shasha DE, Wang JY, Jung JW, Lambert GM, Galbraith DW, Benfey PN. (2003) A gene expression map of the Arabidopsis root. Science 302: 1956–1960 - PubMed
-
- Clough SJ, Bent AF. (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16: 735–743 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

