Looping back to leap forward: transcription enters a new era
- PMID: 24679523
- PMCID: PMC4059561
- DOI: 10.1016/j.cell.2014.02.009
Looping back to leap forward: transcription enters a new era
Abstract
Comparative genome analyses reveal that organismal complexity scales not with gene number but with gene regulation. Recent efforts indicate that the human genome likely contains hundreds of thousands of enhancers, with a typical gene embedded in a milieu of tens of enhancers. Proliferation of cis-regulatory DNAs is accompanied by increased complexity and functional diversification of transcriptional machineries recognizing distal enhancers and core promoters and by the high-order spatial organization of genetic elements. We review progress in unraveling one of the outstanding mysteries of modern biology: the dynamic communication of remote enhancers with target promoters in the specification of cellular identity.
Copyright © 2014 Elsevier Inc. All rights reserved.
Figures

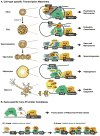
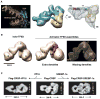
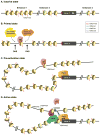

References
-
- Amano T, Sagai T, Tanabe H, Mizushina Y, Nakazawa H, Shiroishi T. Chromosomal dynamics at the Shh locus: limb bud-specific differential regulation of competence and active transcription. Dev Cell. 2009;16:47–57. - PubMed
-
- Andrey G, et al. A switch between topological domains underlies HoxD genes colinearity in mouse limbs. Science. 2013;340:1195–1202. - PubMed
-
- Ang SL, Wierda A, Wong D, Stevens KA, Cascio S, Rossant J, Zaret KS. The formation and maintenance of the definitive endoderm lineage in the mouse: involvement of HNF3/forkhead proteins. Development. 1993;119:1301–1315. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

