The chromatin landscape and transcription factors in T cell programming
- PMID: 24703587
- PMCID: PMC4039984
- DOI: 10.1016/j.it.2014.03.001
The chromatin landscape and transcription factors in T cell programming
Abstract
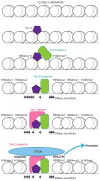
T cell development from multipotent progenitors to specialized effector subsets of mature T cells is guided by the iterative action of transcription factors. At each stage, transcription factors interact not only with an existing landscape of histone modifications and nucleosome packing, but also with other bound factors, while they modify the landscape for later-arriving factors in ways that fundamentally affect the control of gene expression. This review covers insights from genome-wide analyses of transcription factor binding and resulting chromatin conformation changes that reveal roles of cytokine signaling in effector T cell programming, the ways in which one factor can completely transform the impacts of previously bound factors, and the ways in which the baseline chromatin landscape is established during early T cell lineage commitment.
Keywords: CD4(+) T cell subsets; Cis-regulatory element; T cell development; genomics; histone modification.
Copyright © 2014 Elsevier Ltd. All rights reserved.
Conflict of interest statement
The author is not aware of any conflict of interest that could inappropriately influence this work.
Figures

Similar articles
-
The epigenetic landscape of fate decisions in T cells.Nat Immunol. 2025 Apr;26(4):544-556. doi: 10.1038/s41590-025-02113-x. Epub 2025 Mar 19. Nat Immunol. 2025. PMID: 40108419 Review.
-
How transcription factors drive choice of the T cell fate.Nat Rev Immunol. 2021 Mar;21(3):162-176. doi: 10.1038/s41577-020-00426-6. Epub 2020 Sep 11. Nat Rev Immunol. 2021. PMID: 32918063 Free PMC article. Review.
-
Profiling of chromatin accessibility and identification of general cis-regulatory mechanisms that control two ocular lens differentiation pathways.Epigenetics Chromatin. 2019 May 3;12(1):27. doi: 10.1186/s13072-019-0272-y. Epigenetics Chromatin. 2019. PMID: 31053165 Free PMC article.
-
The mouse M-lysozyme gene domain: identification of myeloid and differentiation specific DNasel hypersensitive sites and of a 3'-cis acting regulatory element.Nucleic Acids Res. 1992 Apr 25;20(8):1917-24. doi: 10.1093/nar/20.8.1917. Nucleic Acids Res. 1992. PMID: 1579493 Free PMC article.
-
Divergent functions of hematopoietic transcription factors in lineage priming and differentiation during erythro-megakaryopoiesis.Genome Res. 2014 Dec;24(12):1932-44. doi: 10.1101/gr.164178.113. Epub 2014 Oct 15. Genome Res. 2014. PMID: 25319996 Free PMC article.
Cited by
-
Hepatocellular carcinoma-specific epigenetic checkpoints bidirectionally regulate the antitumor immunity of CD4 + T cells.Cell Mol Immunol. 2024 Nov;21(11):1296-1308. doi: 10.1038/s41423-024-01215-0. Epub 2024 Sep 19. Cell Mol Immunol. 2024. PMID: 39300319
-
Comparative transcriptome analysis of T lymphocyte subpopulations and identification of critical regulators defining porcine thymocyte identity.Front Immunol. 2024 Feb 7;15:1339787. doi: 10.3389/fimmu.2024.1339787. eCollection 2024. Front Immunol. 2024. PMID: 38384475 Free PMC article.
-
Epigenetics of T cell aging.J Leukoc Biol. 2018 Oct;104(4):691-699. doi: 10.1002/JLB.1RI0418-160R. Epub 2018 Jun 27. J Leukoc Biol. 2018. PMID: 29947427 Free PMC article. Review.
-
Transcriptomic features of tumour-infiltrating CD4lowCD8high double positive αβ T cells in melanoma.Sci Rep. 2020 Apr 3;10(1):5900. doi: 10.1038/s41598-020-62664-x. Sci Rep. 2020. PMID: 32246006 Free PMC article.
-
A critical role for c-Myc in group 2 innate lymphoid cell activation.Allergy. 2020 Apr;75(4):841-852. doi: 10.1111/all.14149. Epub 2020 Jan 29. Allergy. 2020. PMID: 31833571 Free PMC article.
References
-
- Welinder E, et al. B-lymphocyte commitment: identifying the point of no return. Semin Immunol. 2011;23:335–340. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

