Analysis of factors that affect FlgM-dependent type III secretion for protein purification with Salmonella enterica serovar Typhimurium
- PMID: 24706743
- PMCID: PMC4054164
- DOI: 10.1128/JB.01572-14
Analysis of factors that affect FlgM-dependent type III secretion for protein purification with Salmonella enterica serovar Typhimurium
Abstract
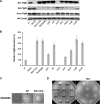
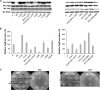
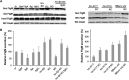
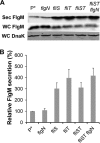
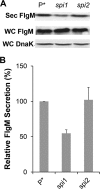
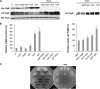
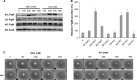
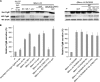
The FlgM protein is secreted in response to flagellar hook-basal body secretion and can be used as a secretion signal to direct selected protein secretion via the flagellar type III secretion (T3S) system [H. M. Singer, M. Erhardt, A. M. Steiner, M. M. Zhang, D. Yoshikami, G. Bulaj, B. M. Olivera, and K. T. Hughes, mBio 3(3):e00115-12, 2012, http://dx.doi.org/10.1128/mBio.00115-12]. Conditions known to affect flagellar gene expression, FlgM stability, and flagellar T3S were tested either alone or in combination to determine their effects on levels of secreted FlgM. These conditions included mutations that affect activity of the flagellar FlhD4C2 master regulatory protein complex or the FlgM T3S chaperone σ(28), the removal of Salmonella pathogenicity island 1 (Spi1), the removal of flagellar late secretion substrates that could compete with FlgM for secretion, and changes in the ionic strength of the growth medium. Conditions that enhanced FlgM secretion were combined in order to maximize levels of secreted FlgM. An optimized FlgM secretion strain was used to secrete and isolate otherwise difficult-to-produce proteins and peptides fused to the C terminus of FlgM. These include cysteine-rich, hydrophobic peptides (conotoxins δ-SVIE and MrVIA), nodule-specific, cysteine-rich antimicrobial peptides (NCR), and a malaria surface antigen domain of apical membrane antigen AMA-1.
Copyright © 2014, American Society for Microbiology. All Rights Reserved.
Figures



Similar articles
-
The flagellar-specific transcription factor, sigma28, is the Type III secretion chaperone for the flagellar-specific anti-sigma28 factor FlgM.Genes Dev. 2006 Aug 15;20(16):2315-26. doi: 10.1101/gad.380406. Genes Dev. 2006. PMID: 16912280 Free PMC article.
-
FlgM is secreted by the flagellar export apparatus in Bacillus subtilis.J Bacteriol. 2015 Jan 1;197(1):81-91. doi: 10.1128/JB.02324-14. Epub 2014 Oct 13. J Bacteriol. 2015. PMID: 25313396 Free PMC article.
-
Flk prevents premature secretion of the anti-sigma factor FlgM into the periplasm.Mol Microbiol. 2006 May;60(3):630-43. doi: 10.1111/j.1365-2958.2006.05135.x. Mol Microbiol. 2006. PMID: 16629666 Free PMC article.
-
Coupling of flagellar gene expression to flagellar assembly in Salmonella enterica serovar typhimurium and Escherichia coli.Microbiol Mol Biol Rev. 2000 Dec;64(4):694-708. doi: 10.1128/MMBR.64.4.694-708.2000. Microbiol Mol Biol Rev. 2000. PMID: 11104815 Free PMC article. Review.
-
Regulation of flagellar assembly.Curr Opin Microbiol. 2002 Apr;5(2):160-5. doi: 10.1016/s1369-5274(02)00302-8. Curr Opin Microbiol. 2002. PMID: 11934612 Review.
Cited by
-
Extracellular matrix-associated proteins form an integral and dynamic system during Pseudomonas aeruginosa biofilm development.Front Cell Infect Microbiol. 2015 May 13;5:40. doi: 10.3389/fcimb.2015.00040. eCollection 2015. Front Cell Infect Microbiol. 2015. PMID: 26029669 Free PMC article.
-
Type III secretion systems: the bacterial flagellum and the injectisome.Philos Trans R Soc Lond B Biol Sci. 2015 Oct 5;370(1679):20150020. doi: 10.1098/rstb.2015.0020. Philos Trans R Soc Lond B Biol Sci. 2015. PMID: 26370933 Free PMC article. Review.
-
Controlling tumor progression and recurrence in mice through combined treatment with a PD-L1 inhibitor and a designer Salmonella strain that delivers GM-CSF.Acta Pharm Sin B. 2024 Dec;14(12):5479-5492. doi: 10.1016/j.apsb.2024.07.011. Epub 2024 Jul 10. Acta Pharm Sin B. 2024. PMID: 39807328 Free PMC article.
-
Inefficient Secretion of Anti-sigma Factor FlgM Inhibits Bacterial Motility at High Temperature.iScience. 2019 Jun 28;16:145-154. doi: 10.1016/j.isci.2019.05.022. Epub 2019 May 20. iScience. 2019. PMID: 31170626 Free PMC article.
-
Bacterial type III secretion systems: a complex device for the delivery of bacterial effector proteins into eukaryotic host cells.FEMS Microbiol Lett. 2018 Oct 1;365(19):fny201. doi: 10.1093/femsle/fny201. FEMS Microbiol Lett. 2018. PMID: 30107569 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

