Regulation of microtubule minus-end dynamics by CAMSAPs and Patronin
- PMID: 24706919
- PMCID: PMC4000804
- DOI: 10.1073/pnas.1404133111
Regulation of microtubule minus-end dynamics by CAMSAPs and Patronin
Abstract
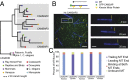
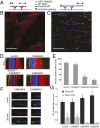
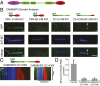
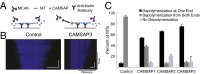
The microtubule (MT) cytoskeleton plays an essential role in mitosis, intracellular transport, cell shape, and cell migration. The assembly and disassembly of MTs, which can occur through the addition or loss of subunits at the plus- or minus-ends of the polymer, is essential for MTs to carry out their biological functions. A variety of proteins act on MT ends to regulate their dynamics, including a recently described family of MT minus-end binding proteins called calmodulin-regulated spectrin-associated protein (CAMSAP)/Patronin/Nezha. Patronin, the single member of this family in Drosophila, was previously shown to stabilize MT minus-ends against depolymerization in vitro and in vivo. Here, we show that all three mammalian CAMSAP family members also bind specifically to MT minus-ends and protect them against kinesin-13-induced depolymerization. However, these proteins differ in their abilities to suppress tubulin addition at minus-ends and to dissociate from MTs. CAMSAP1 does not interfere with polymerization and tracks along growing minus-ends. CAMSAP2 and CAMSAP3 decrease the rate of tubulin incorporation and remain bound, thereby creating stretches of decorated MT minus-ends. By using truncation analysis, we find that somewhat different minimal domains of CAMSAP and Patronin are involved in minus-end localization. However, we find that, in both cases, a highly conserved C-terminal domain and a more variable central domain cooperate to suppress minus-end dynamics in vitro and that both regions are required to stabilize minus-ends in Drosophila S2 cells. These results show that members of the CAMSAP/Patronin family all localize to and protect minus-ends but have evolved distinct effects on MT dynamics.
Keywords: TIRF microscopy; cytoskeletal regulation; tubulin polymerization.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

References
-
- Mitchison TJ. Localization of an exchangeable GTP binding site at the plus end of microtubules. Science. 1993;261(5124):1044–1047. - PubMed
-
- Mitchison T, Kirschner M. Dynamic instability of microtubule growth. Nature. 1984;312(5991):237–242. - PubMed
-
- Bieling P, et al. Reconstitution of a microtubule plus-end tracking system in vitro. Nature. 2007;450(7172):1100–1105. - PubMed
-
- Akhmanova A, Steinmetz MO. Tracking the ends: A dynamic protein network controls the fate of microtubule tips. Nat Rev Mol Cell Biol. 2008;9(4):309–322. - PubMed
-
- Schuyler SC, Pellman D. Microtubule “plus-end-tracking proteins”: The end is just the beginning. Cell. 2001;105(4):421–424. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

