Gap-directed translesion DNA synthesis of an abasic site on circular DNA templates by a human replication complex
- PMID: 24710081
- PMCID: PMC3977967
- DOI: 10.1371/journal.pone.0093908
Gap-directed translesion DNA synthesis of an abasic site on circular DNA templates by a human replication complex
Abstract
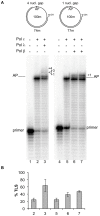
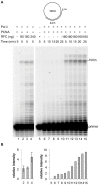
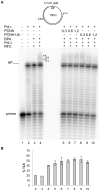
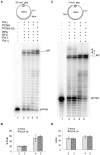
DNA polymerase ε (pol ε) is believed to be the leading strand replicase in eukaryotes whereas pols λ and β are thought to be mainly involved in re-synthesis steps of DNA repair. DNA elongation by the human pol ε is halted by an abasic site (apurinic/apyrimidinic (AP) site). We have previously reported that human pols λ, β and η can perform translesion synthesis (TLS) of an AP site in the presence of pol ε. In the case of pol λ and β, this TLS requires the presence of a gap downstream from the product synthetized by the ε replicase. However, since these studies were conducted exclusively with a linear DNA template, we decided to test whether the structure of the template could influence the capacity of the pols ε, λ, β and η to perform TLS of an AP site. Therefore, we have investigated the replication of damaged "minicircle" DNA templates. In addition, replication of circular DNA requires, beyond DNA pols, the processivity clamp PCNA, the clamp loader replication factor C (RFC), and the accessory proteins replication protein A (RPA). Finally we have compared the capacity of unmodified versus monoubiquitinated PCNA in sustaining TLS by pols λ and η on a circular template. Our results indicate that in vitro gap-directed TLS synthesis by pols λ and β in the presence of pol ε, RPA and PCNA is unaffected by the structure of the DNA template. Moreover, monoubiquitination of PCNA does not affect TLS by pol λ while it appears to slightly stimulate TLS by pol η.
Conflict of interest statement
Figures


Similar articles
-
In vitro gap-directed translesion DNA synthesis of an abasic site involving human DNA polymerases epsilon, lambda, and beta.J Biol Chem. 2011 Sep 16;286(37):32094-104. doi: 10.1074/jbc.M111.246611. Epub 2011 Jul 13. J Biol Chem. 2011. PMID: 21757740 Free PMC article.
-
DNA polymerase δ-interacting protein 2 is a processivity factor for DNA polymerase λ during 8-oxo-7,8-dihydroguanine bypass.Proc Natl Acad Sci U S A. 2013 Nov 19;110(47):18850-5. doi: 10.1073/pnas.1308760110. Epub 2013 Nov 4. Proc Natl Acad Sci U S A. 2013. PMID: 24191025 Free PMC article.
-
Translesion synthesis across abasic lesions by human B-family and Y-family DNA polymerases α, δ, η, ι, κ, and REV1.J Mol Biol. 2010 Nov 19;404(1):34-44. doi: 10.1016/j.jmb.2010.09.015. Epub 2010 Oct 1. J Mol Biol. 2010. PMID: 20888339 Free PMC article.
-
A Comprehensive View of Translesion Synthesis in Escherichia coli.Microbiol Mol Biol Rev. 2020 Jun 17;84(3):e00002-20. doi: 10.1128/MMBR.00002-20. Print 2020 Aug 19. Microbiol Mol Biol Rev. 2020. PMID: 32554755 Free PMC article. Review.
-
Translesion synthesis in Escherichia coli: lessons from the NarI mutation hot spot.DNA Repair (Amst). 2007 Jul 1;6(7):1032-41. doi: 10.1016/j.dnarep.2007.02.021. Epub 2007 Apr 2. DNA Repair (Amst). 2007. PMID: 17403618 Review.
Cited by
-
EGFP Reporters for Direct and Sensitive Detection of Mutagenic Bypass of DNA Lesions.Biomolecules. 2020 Jun 13;10(6):902. doi: 10.3390/biom10060902. Biomolecules. 2020. PMID: 32545792 Free PMC article.
-
Translesion DNA polymerases in eukaryotes: what makes them tick?Crit Rev Biochem Mol Biol. 2017 Jun;52(3):274-303. doi: 10.1080/10409238.2017.1291576. Epub 2017 Mar 9. Crit Rev Biochem Mol Biol. 2017. PMID: 28279077 Free PMC article. Review.
References
-
- Kornberg A, Baker TA (2005) DNA Replication. 2nd ed. University Science Books.
-
- Hübscher U, Spadari S, Villani G, Maga G (2010) DNA Polymerases: Discovery, Characterization and Functions in Cellular DNA Transactions. 1st ed. World Scientific Publishing Company.
-
- Lindahl T (1993) Instability and decay of the primary structure of DNA. Nature 362: 709–715. - PubMed
-
- Friedberg EC, Walker GC, Siede W, Wood RD, Schultz RA, et al... (2005) DNA Repair And Mutagenesis. 2nd ed. ASM Press.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

